Get isoTar
isoTar is a high performance web-based containerized application for Consensus miRNA Targeting Prediction and Functional Enrichment Analysis. It leverages on Docker, a virtual environment manager, independent of the host Operating System (OS). Through a friendly web-based User Interface (UI), users can explore the impact of Human miRNA variants on the Human Transcriptome.
Disclaimer: This program is for educational and research purposes only.
This program is free software; you can redistribute it and/or modify
it under the terms of the GNU General Public License as published by
the Free Software Foundation; either version 3 of the License, or (at
your option) any later version.
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU
General Public License for more details.
A copy of the license text can be obtained from https://www.gnu.org/licenses/gpl-3.0.en.html
Available for
MD5: 8b85f24f197e0c7efbb08ac5431f4484
SHA1: 1ad1ac20497f15caddf96cfa77f90eb687ddf12c
SHA256: 1a8d6fc72a8308791aeb731138d9e7d7e3fdc0fb075f204abf30cdaad31a3ea1
How to cite
Distefano R., Nigita G., Veneziano D., Romano G., Croce CM., Acunzo M. "isoTar: Consensus Target Prediction with Enrichment Analysis for MicroRNAs Harboring Editing Sites and Other Variations." Methods Mol Biol. (2019)
PubMedOverview
Powered by
Resources included in isoTar
Prediction tools included in isoTar
Installation
isoTar requires Docker
installed on your PC. Please follow the instructions associated to your Operating System (Ubuntu (Linux),
Mac, or
Windows). You can find
instructions for additional Platforms/Operating Systems here.
Once you get Docker, you can proceed with the isoTar installation as specified below:
1. Download isoTar (click on the button above).
2. Decompress the Zip file.
3. Open Terminal (Linux/Mac), Powershell (Windows).
4. Install isoTar (image name: isotar_v1.2.1.tar) using the following command:
docker load -i PATH_TO_THE_TAR_FILE/isotar_v1.2.1.tarThe installation process will generate an output similar to the one reported below:
34394faab172: Loading layer [==================================================>] 17.92kB/17.92kB
fbf57c7a838c: Loading layer [==================================================>] 11.78kB/11.78kB
2d63176664ff: Loading layer [==================================================>] 5.632kB/5.632kB
5ab785ed50da: Loading layer [==================================================>] 3.072kB/3.072kB
097adadfb283: Loading layer [==================================================>] 7.429GB/7.429GB
86c87be6face: Loading layer [==================================================>] 108.6MB/108.6MB
2dcd42fb11cf: Loading layer [==================================================>] 720.3MB/720.3MB
a64e27523e49: Loading layer [==================================================>] 103.6MB/103.6MB
1975cd33bc84: Loading layer [==================================================>] 1.536kB/1.536kB
e518cf78e956: Loading layer [==================================================>] 77.04MB/77.04MB
890769017077: Loading layer [==================================================>] 817.8MB/817.8MB
8085b1d466a4: Loading layer [==================================================>] 84.26MB/84.26MB
42d84f02a1ed: Loading layer [==================================================>] 1.778MB/1.778MB
c01029da65f8: Loading layer [==================================================>] 26.83MB/26.83MB
e6aaf08a8d15: Loading layer [==================================================>] 238.7MB/238.7MB
7ec2c40ad70d: Loading layer [==================================================>] 3.402MB/3.402MB
2890d18c1855: Loading layer [==================================================>] 784.9kB/784.9kB
fff38d1ba0f3: Loading layer [==================================================>] 6.212MB/6.212MB
e854dde956f4: Loading layer [==================================================>] 4.008MB/4.008MB
0dc1195a50cd: Loading layer [==================================================>] 156.6MB/156.6MB
17f94a00c15c: Loading layer [==================================================>] 337.4MB/337.4MB
f1699f58bd58: Loading layer [==================================================>] 2.048kB/2.048kB
dcc74bbe2463: Loading layer [==================================================>] 3.072kB/3.072kB
9eb14f476dd0: Loading layer [==================================================>] 3.584kB/3.584kB
30ff6d340466: Loading layer [==================================================>] 3.072kB/3.072kB
aa61ab025cdc: Loading layer [==================================================>] 4.096kB/4.096kB
8494dd294d5f: Loading layer [==================================================>] 4.096kB/4.096kB
0dc19cc08570: Loading layer [==================================================>] 3.584kB/3.584kB
90acf0e9a3a3: Loading layer [==================================================>] 2.56kB/2.56kB
416dc1488da0: Loading layer [==================================================>] 2.56kB/2.56kB
ec4f7b5e60cb: Loading layer [==================================================>] 2.56kB/2.56kB
21adbf10d522: Loading layer [==================================================>] 2.56kB/2.56kB
46acdfccfc0a: Loading layer [==================================================>] 4.096kB/4.096kB
Loaded image: isotar:v1.2.1The previous command will install the isoTar image on your PC. By running
docker imagesyou will list Docker images available on your PC. You can verify if the installation process has been executed correctly by checking if the isotar image is showed as reported below:
REPOSITORY TAG IMAGE ID CREATED SIZE
isotar v1.2.1 15e0f7fa60fe 44 minutes ago 10.4GBKeep in mind that the IMAGE ID showed here represents an example, since you will see a different one on your PC.
4. Run isoTar using the following command:
$ docker run --rm --name isotar -d -p 80:8080 isotar:v1.2.1
The previous command will run isoTar as a Web Service on your PC. In particular, it will bind the 80 port of your PC with the isoTar Web Service 8080 port.
You can specify a different port for your PC if it is already used by other services. For example, you could use port 8080 or similar instead of 80 (docker run --rm --name isotar -d -p 8080:8080 isotar:v1.2.1).
Note This command will create an instance of the isoTar image previously installed on your PC. As consequence, you will be required to run such command every time you will run/reboot
your PC, whenever you need to use isoTar.
5. Open your web browser and write the URL localhost, if you used port 80 on your PC. If you specified a different port, for example 8080 port, use the URL localhost:8080
How to
Consensus Prediction + Functional Analysis, as well as Functional Enrichment Analysis only input setups are available through the corresponding button.
Input setup
Consensus Prediction + Functional Analysis Functional Enrichment Analysis only
STEP 1 miRNA(s) setup
There are three ways to load a list of miRNA(s) and their variation(s):
- Using the "Search and customize a miRNA" form (
[Option 1]) - Using the "Known Variations Databases" button (
[Option 2]) - Pasting them directly into the "textarea" (
[Option 3])
These options are shown in the following figure:
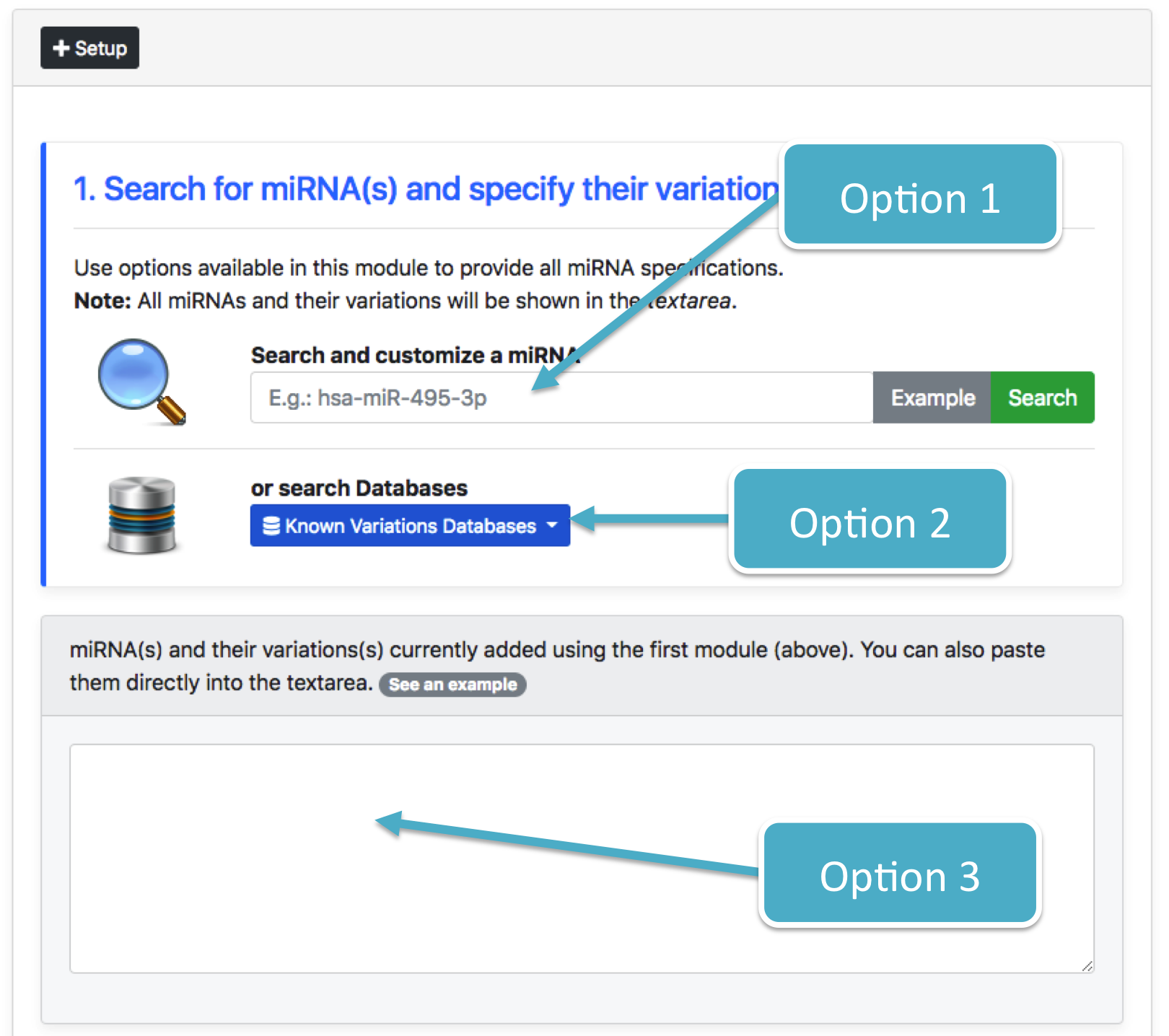
Each option requires specific steps in order to generate the correct input format for the analysis. Whatever option is chosen, each miRNA specification must be compliant with the isoTar format (see below).
>hsa-miR-XXX hsa-[mir/let]-XXX <variations list>
mature miRNA sequence
>hsa-let-7c-5p hsa-let-7c 17:A|G;+1|-2
UGAGGUAGUAGGUUGUAUGGUU
hsa-let-7c-5p, hsa-let-7c, 17:A|G;+1|-2, and UGAGGUAGUAGGUUGUAUGGUU represent, respectively,
the mature miRNA name (Human only), the pre-miRNA name (Human only), the variations list (in this example, a nucleotide and an isomiR modification),
and the mature miRNA reference sequence (Human only).
[Option 1] Search and customize a miRNA
Using this module, it is possible to search for a specific mature miRNA. It shows a list of miRNA(s) and their pre-miRNA(s).
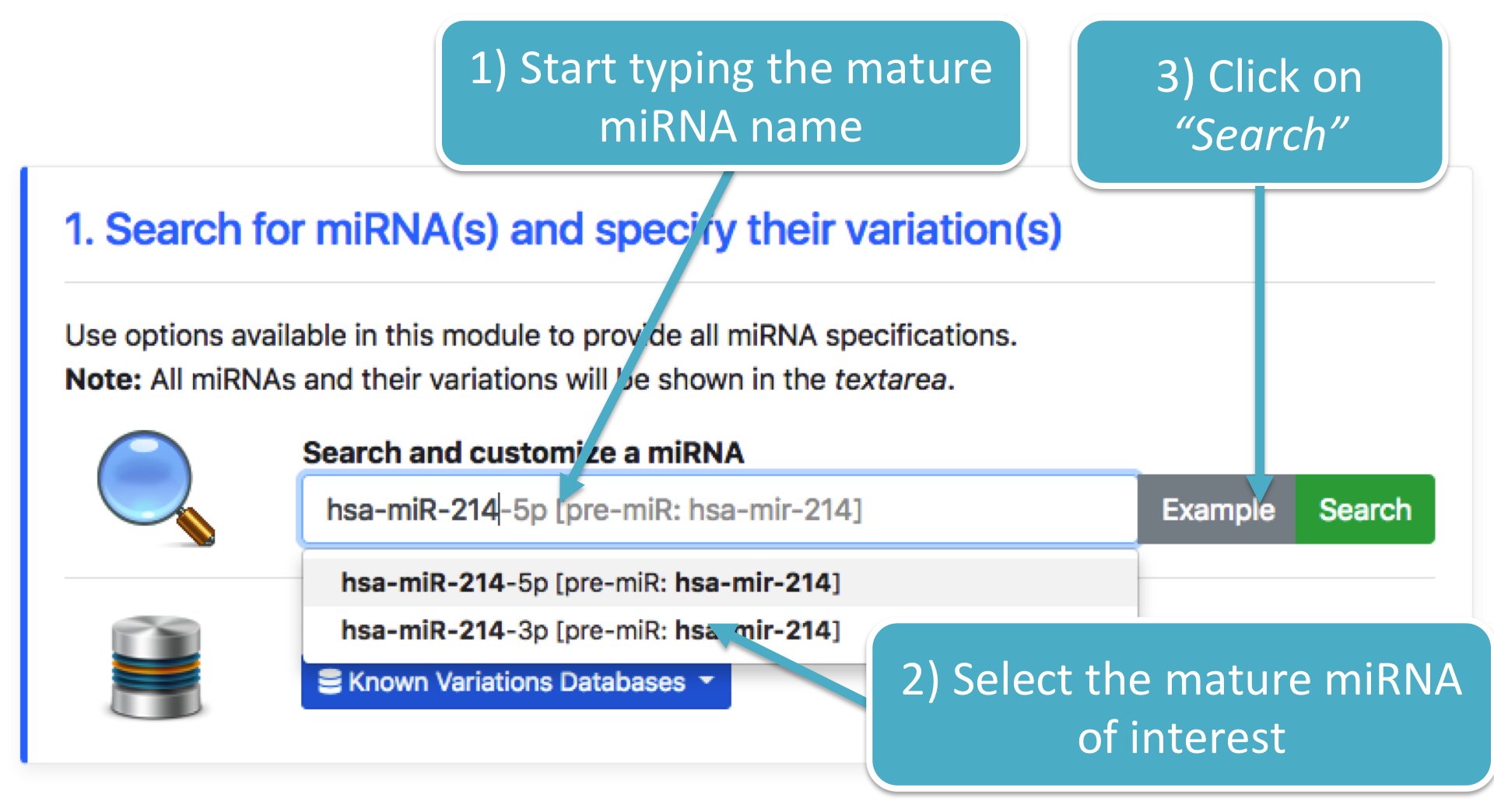

Once a miRNA has been selected, click on the Search button in order to open the customization module (figure below).
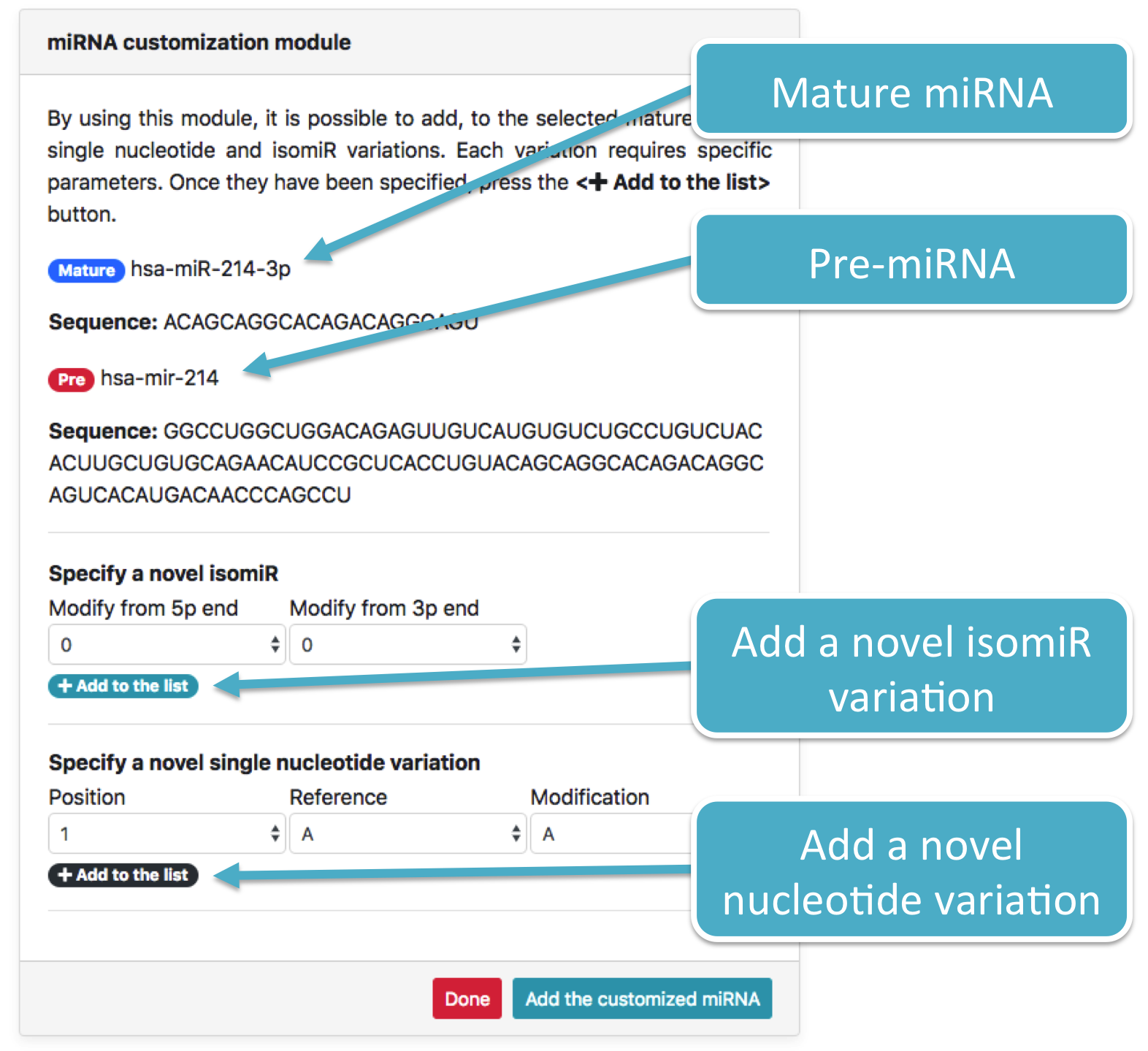
Two distinct variations are shown in the following figure:
-
A single nucleotide variation at position
6that changes the nucleotide fromA(Ref) toG(Mod) -
A single isomiR variation that adds one nucleotide at
5pend, resulting in an extension of the miRNA sequence. Added nucleotides are retrieved from the pre-miRNA which the selected miRNA originates from.
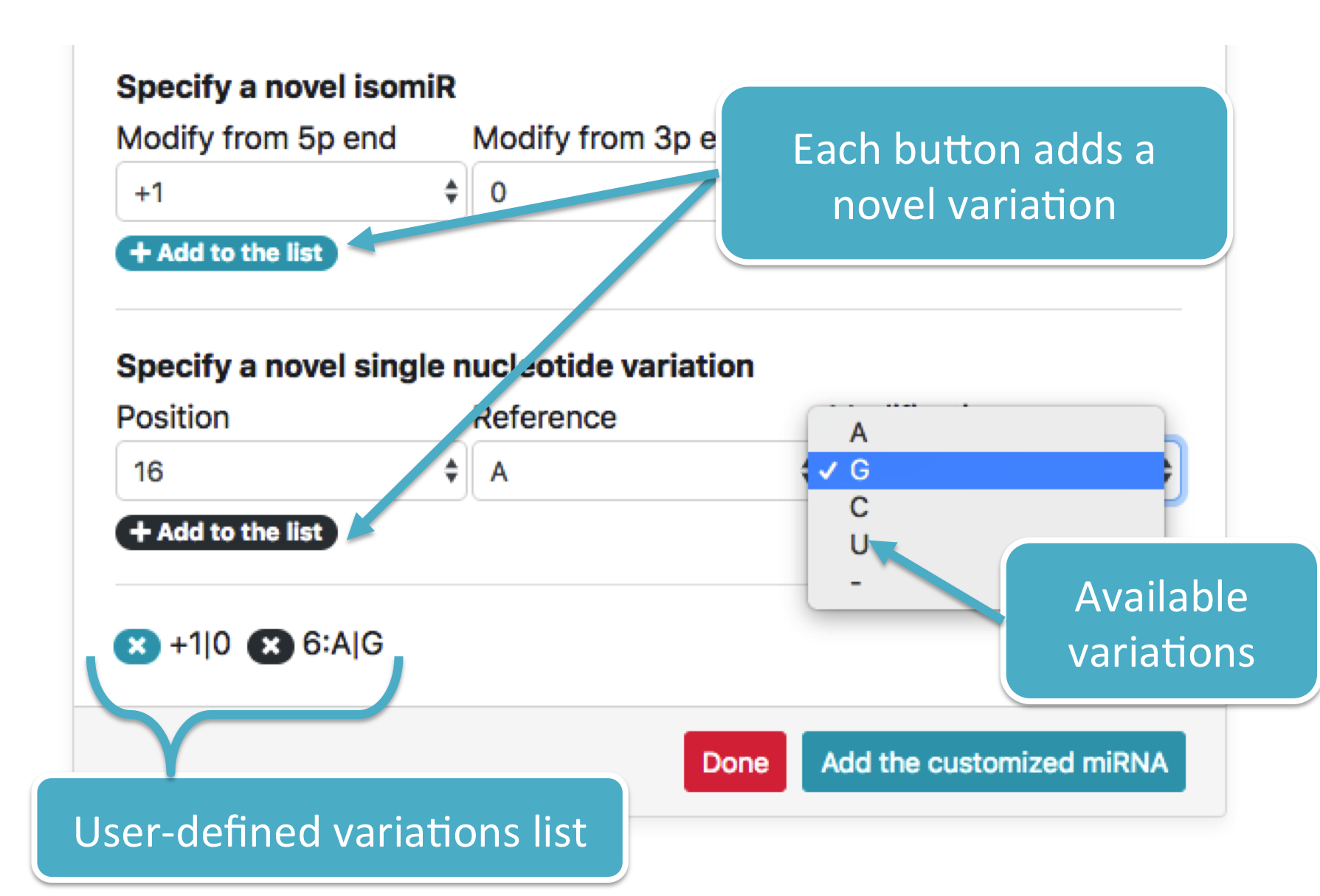
Once finished, press the <Add customized miRNA> button to add all variations to the textarea. The textarea will look like this:
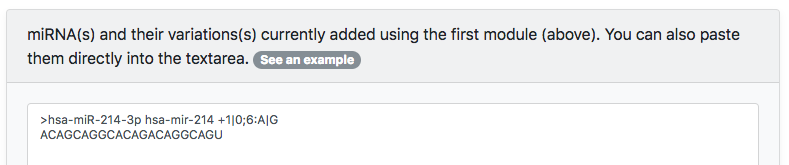
[Option 2] Known Variations Databases
An alternative way to specify miRNA variations is to select them using one or more of the databases provided with isoTar.
These databases can be selected using the following dropdown menu (figure below).
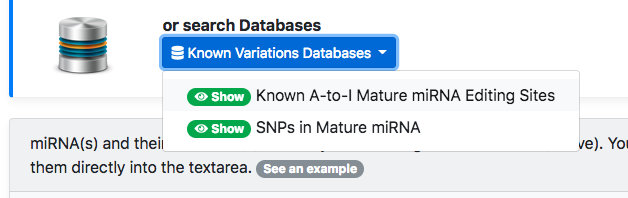
Each database can be consulted through a user-friendly interface which lists variations in a tabular form.
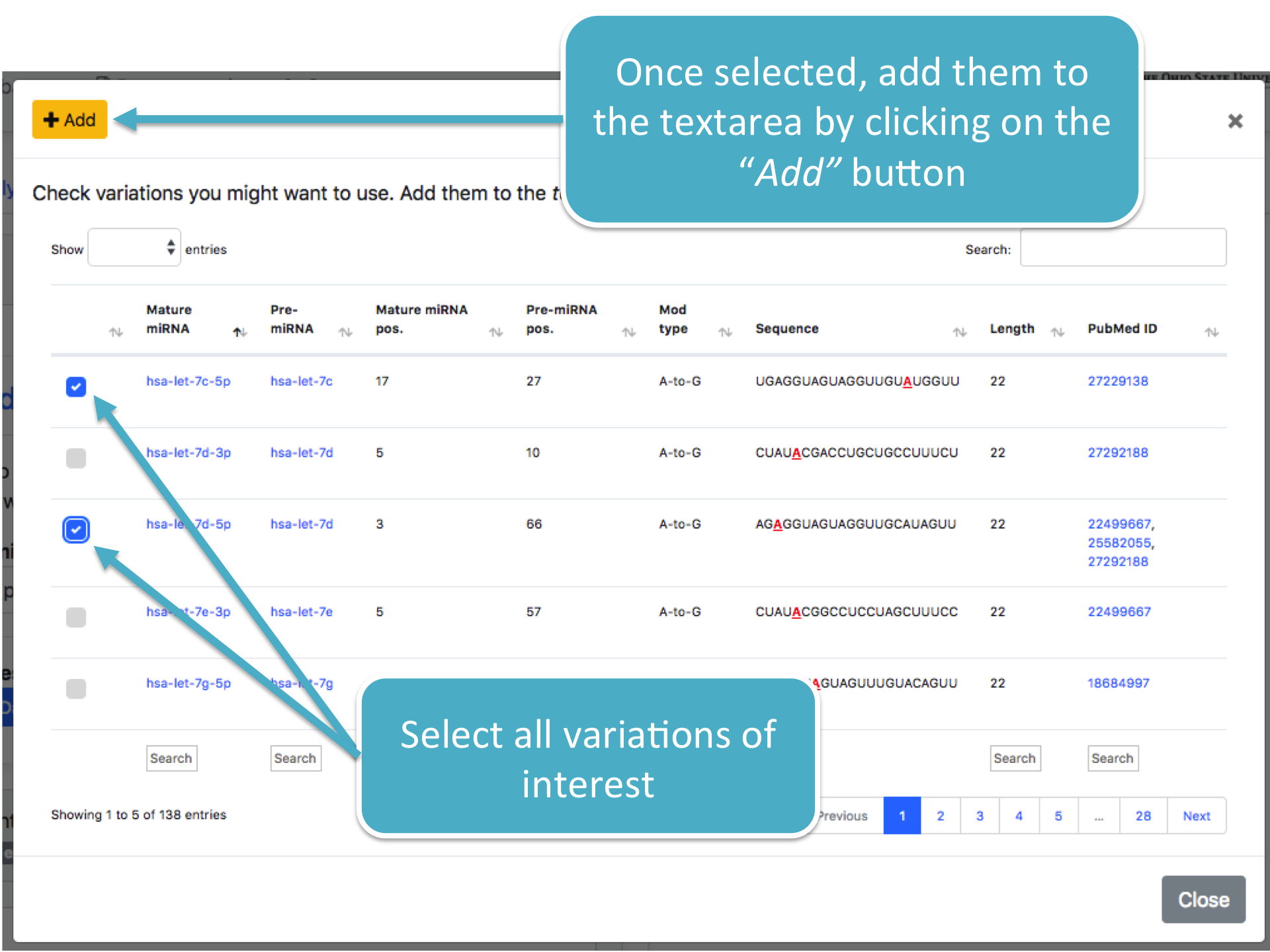
STEP 2 Tools selection and Minimum Consensus
Before performing any prediction analysis, it is necessary to select which tools should be executed and, possibly, their combination by specifying the AND/OR option (if one or more tools must/can identify the same predicted target). In addition, a Minimum Consensus has to be provided as well. It corresponds to the minimum number of tools that must or can identify the same predicted target. The following figure shows these notions:
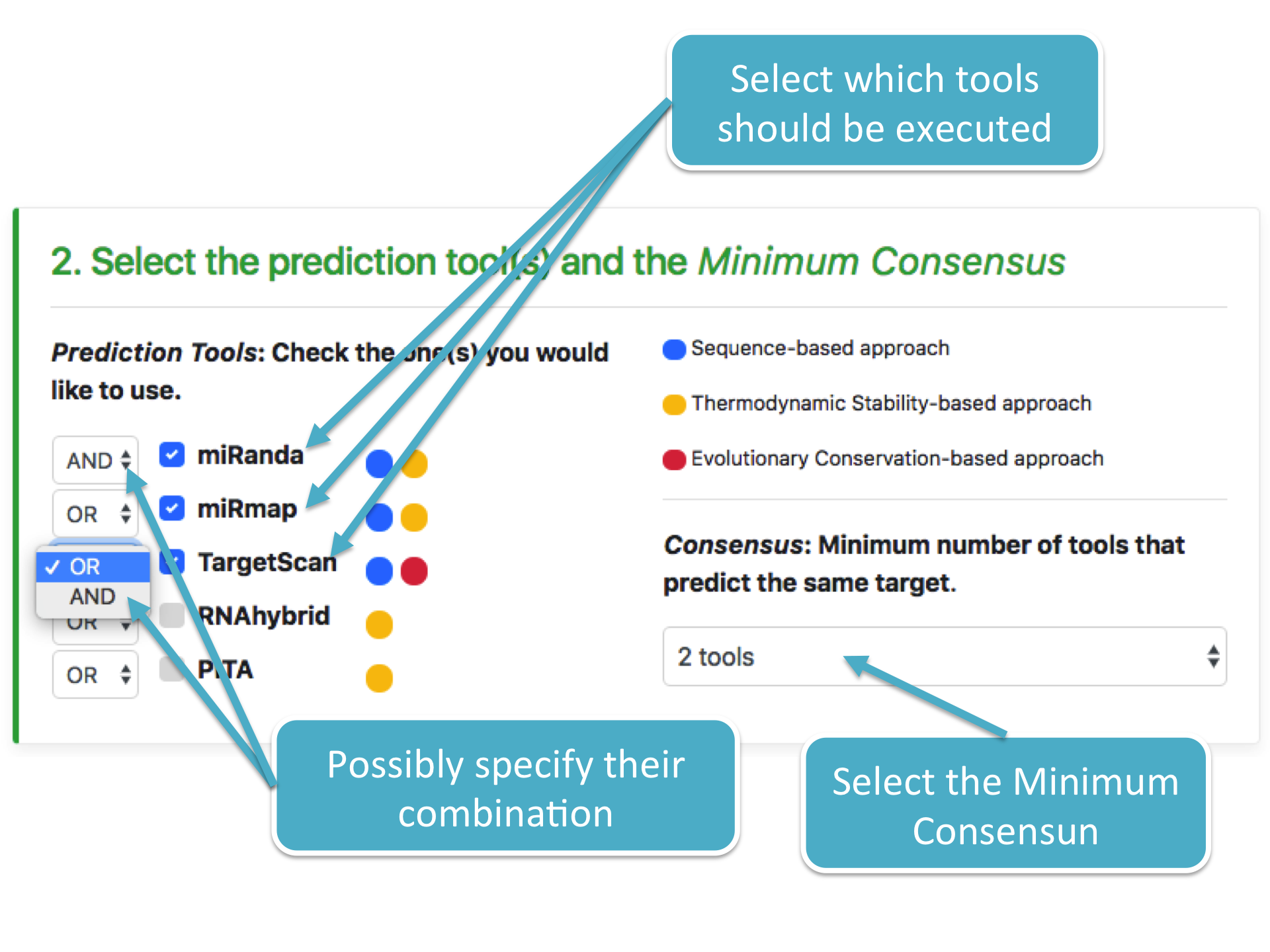
STEP 3 CPU cores selection
It is possible to perform all the analyses using a single core CPU or taking advantage of a multiple CPU cores. In this latter case, tools and the analysis
are executed in parallel in order to optimize execution time, hiding the underlying complexity to the user.
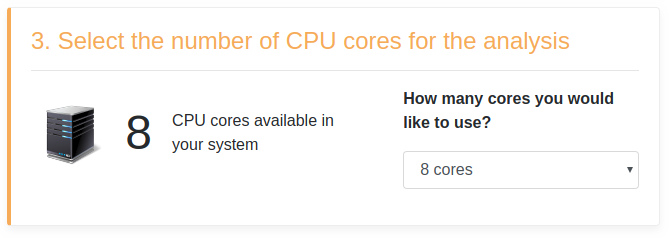
The following figure shows a hypothetical system equipped with 8 CPU cores, as well as a dropdown menu reporting all selectable CPU cores.
In this example, it is possible to select from 1 to 8 cores.
Consensus Prediction input submission & analysis execution
Click on the Submit button to start input verification (the button is located below the STEP 3 box). Then, isoTar will show you a summary of all submitted miRNA(s) and their variation(s). Results are grouped according to the specified variation types, such as single nucleotide variation, isomiR, and/or wild type miRNA (no variation specified). The figure below shows variations grouped for Single nucleotide variation and isomiR.
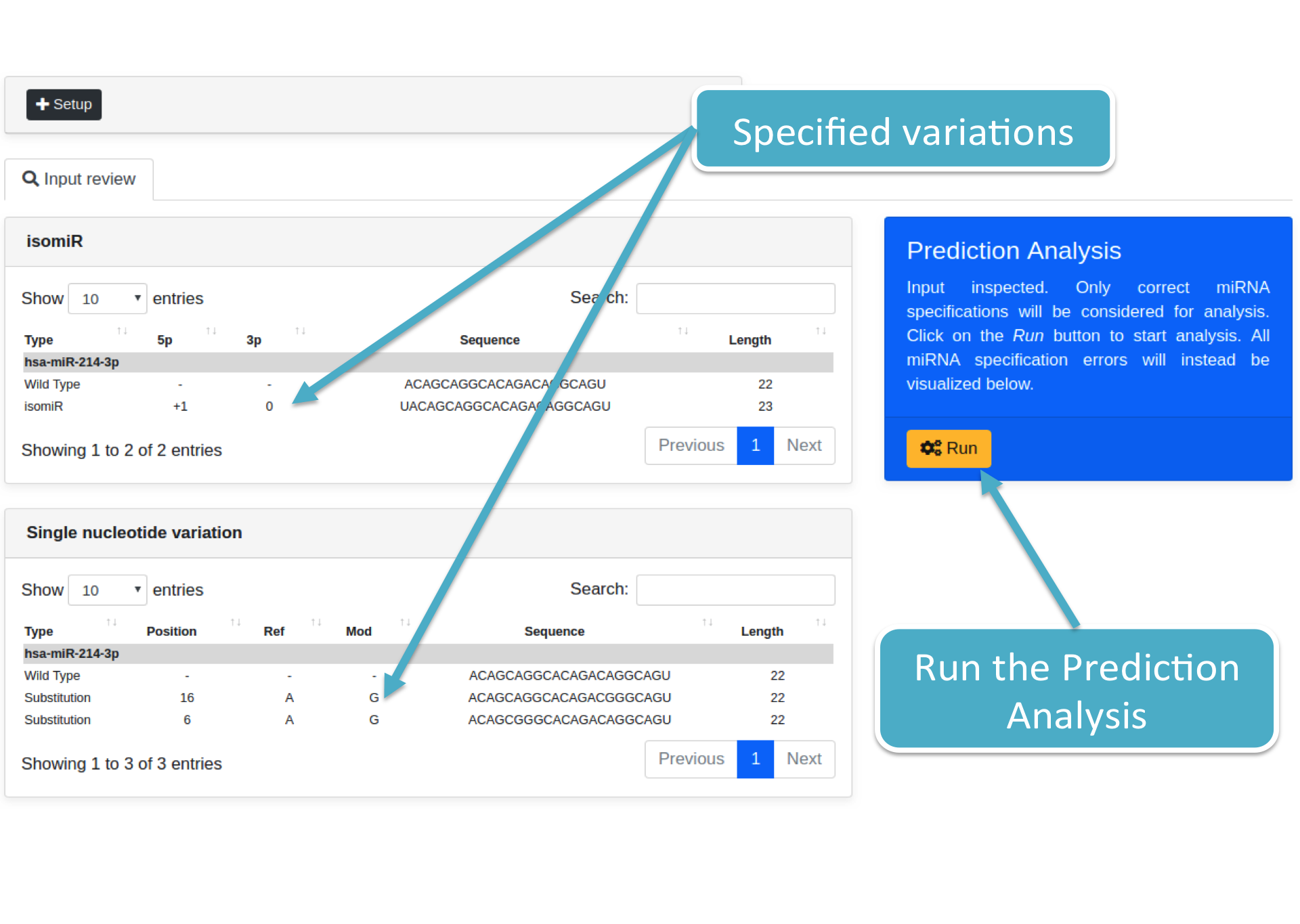
Once you reviewed your input, click on the Run button to start the Prediction Analysis. isoTar will show you a box reporting the analysis status, as well as the stopping button to shut down its execution (both shown below).
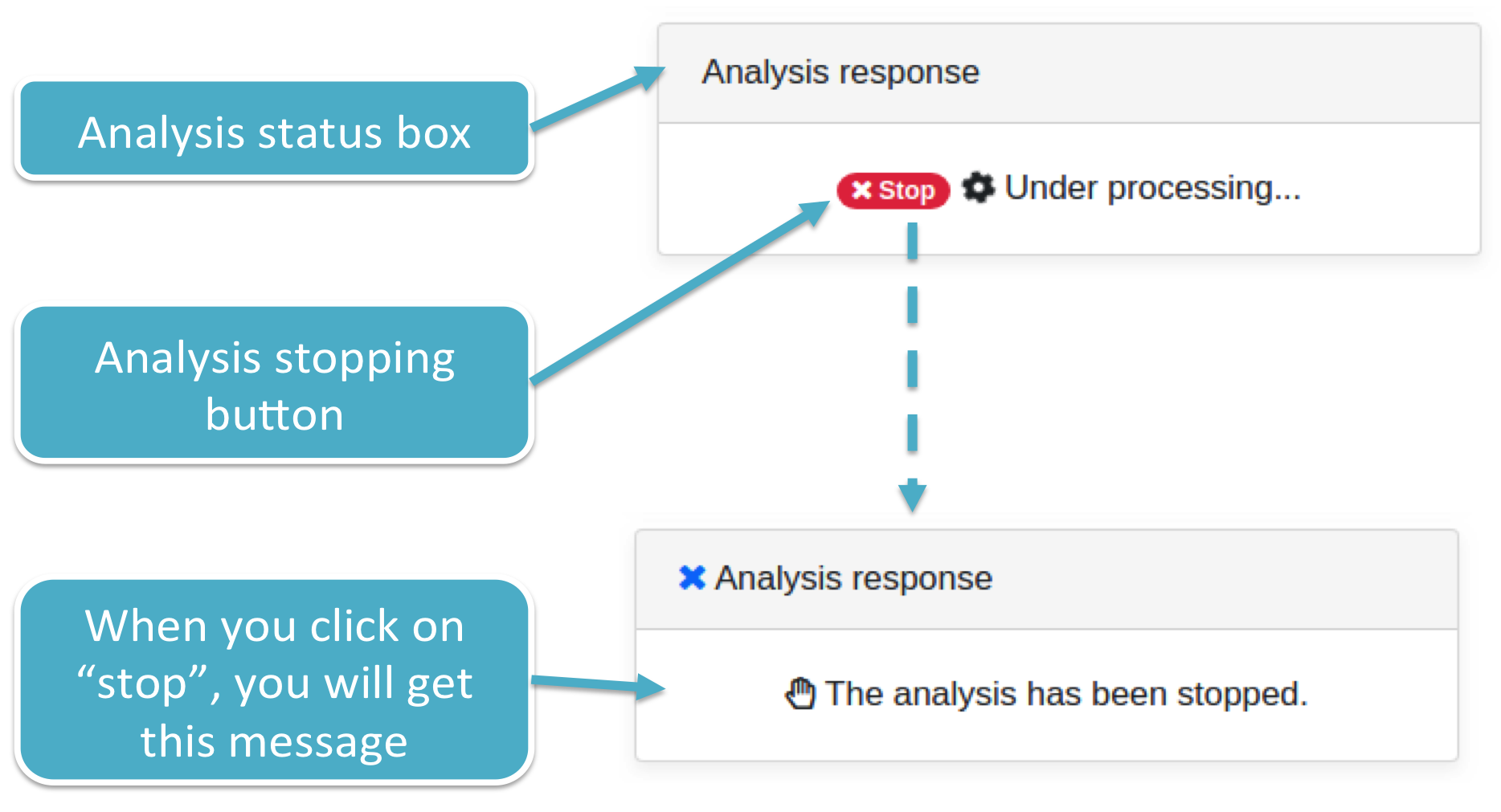
STEP 1 P-value specifics
Results generated through the Functional Analysis are filtered according to the user-specified P-value:
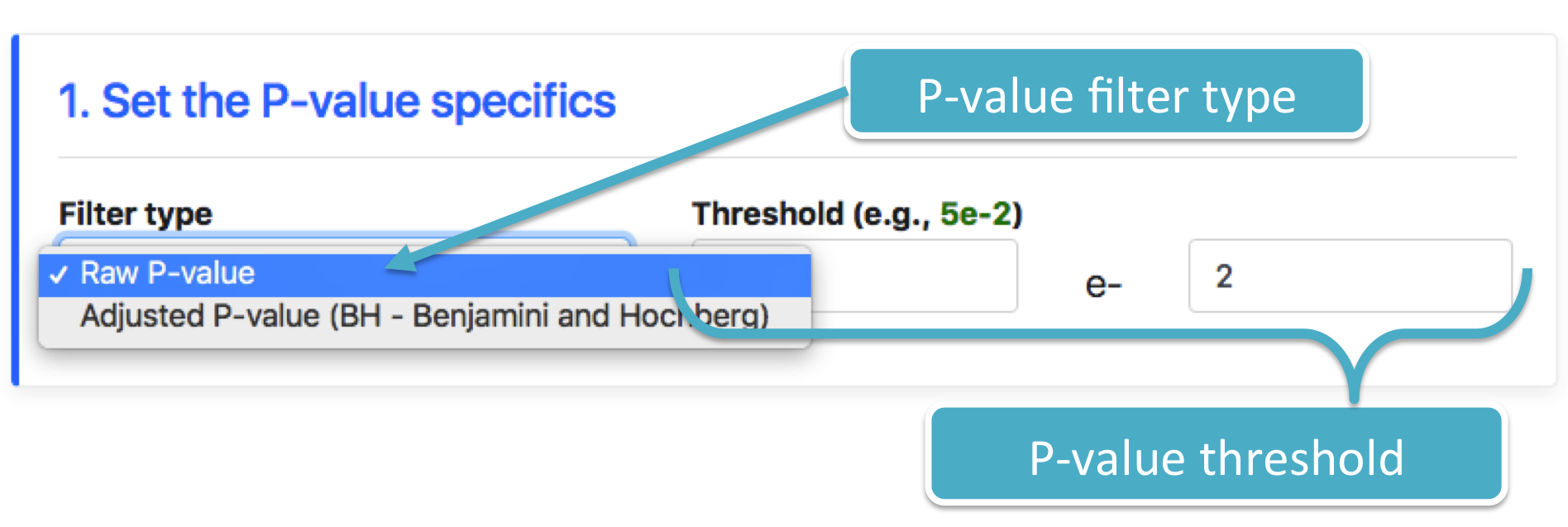
- Type, which represents the type of the P-value that must be taken into account in the filtering. A user can choose to filter on: (Raw P-value)-based filter or (Adjusted P-value)-based filter (Benjamini and Hochberg).
- Threshold, which represents the upper bound limit value for the set of P-values the user wishes to display.
The figure below shows the module which allows to set the P-value type and threshold.
STEP 2 Resources selection
The Functional Analysis can be performed using different resources:
Select the resources in the Functional Analysis to be considered, as shown in the following figure:

STEP 3 CPU cores selection
It is possible to perform all the analyses using a single core CPU or taking advantage of a multiple CPU cores. In this latter case, tools and the analysis
are executed in parallel in order to optimize execution time, hiding the underlying complexity to the user.
The following figure shows a hypothetical system equipped with 8 CPU cores, as well as a dropdown menu reporting all selectable CPU cores.
In this example, it is possible to select from 1 to 8 cores.

Analysis execution
Click on the Submit button to start the Functional Analysis. isoTar will show you a box reporting the analysis status, as well as the stopping button to shut down its execution (both shown below).

STEP 1 Gene list(s)
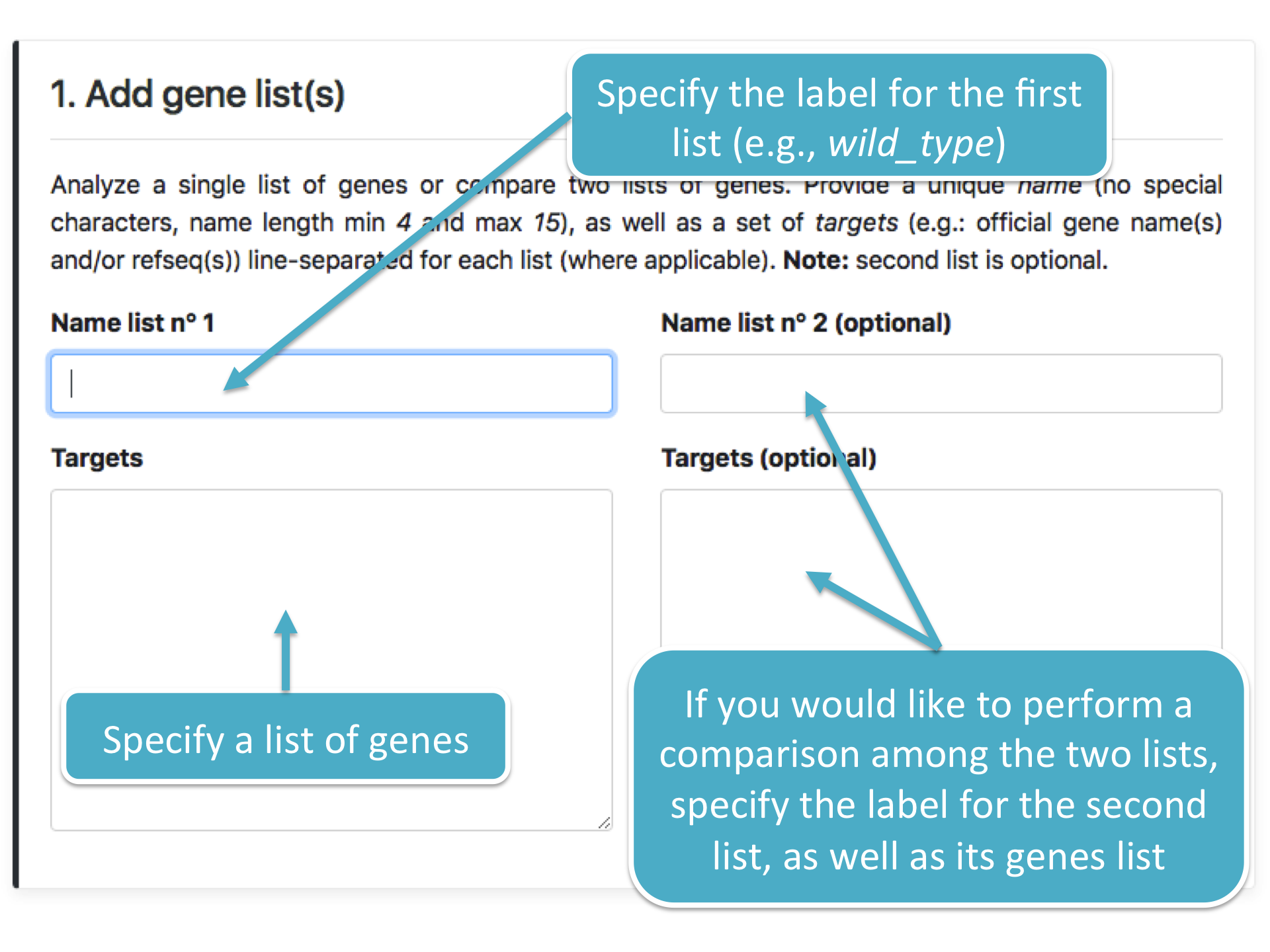
This module (the figure below) allows to specify whether the Functional Enrichment Analysis must be performed on one or two lists of genes, by specifying their
official gene name (e.g.: PTEN) and/or refseq ID (e.g.: NM_000314).
If two lists are specified, the analysis is performed by comparing both lists. As an example, a user could be interested in comparing two lists of genes, each one associated with a specific
biological condition (e.g.: Wild Type vs. Knockout).
Both the first label and the list of genes are mandatory. However, it is not necessary to specify the second set of label and genes list if no comparison must be performed.
STEP 2 P-value specifics
Results generated through the Functional Analysis are filtered according to the user-specified P-value:
- Type, which represents the type of the P-value that must be taken into account in the filtering. A user can choose to filter on: (Raw P-value)-based filter or (Adjusted P-value)-based filter (Benjamini and Hochberg).
- Threshold, which represents the upper bound limit value for the set of P-values the user wishes to display.
The figure below shows the module which allows to set the P-value type and threshold.

STEP 3 Resources selection
The Functional Analysis can be performed using different resources:
Select the resources in the Functional Analysis to be considered, as shown in the following figure:

STEP 4 CPU cores selection
It is possible to perform all the analyses using a single core CPU or taking advantage of a multiple CPU cores. In this latter case, tools and the analysis
are executed in parallel in order to optimize execution time, hiding the underlying complexity to the user.
The following figure shows a hypothetical system equipped with 8 CPU cores, as well as a dropdown menu reporting all selectable CPU cores.
In this example, it is possible to select from 1 to 8 cores.

Results explanation
Consensus Prediction + Functional Analysis Functional Enrichment Analysis only
After the conclusion of the analysis, prediction results will be shown in a new tab panel (named Results), as shown below:
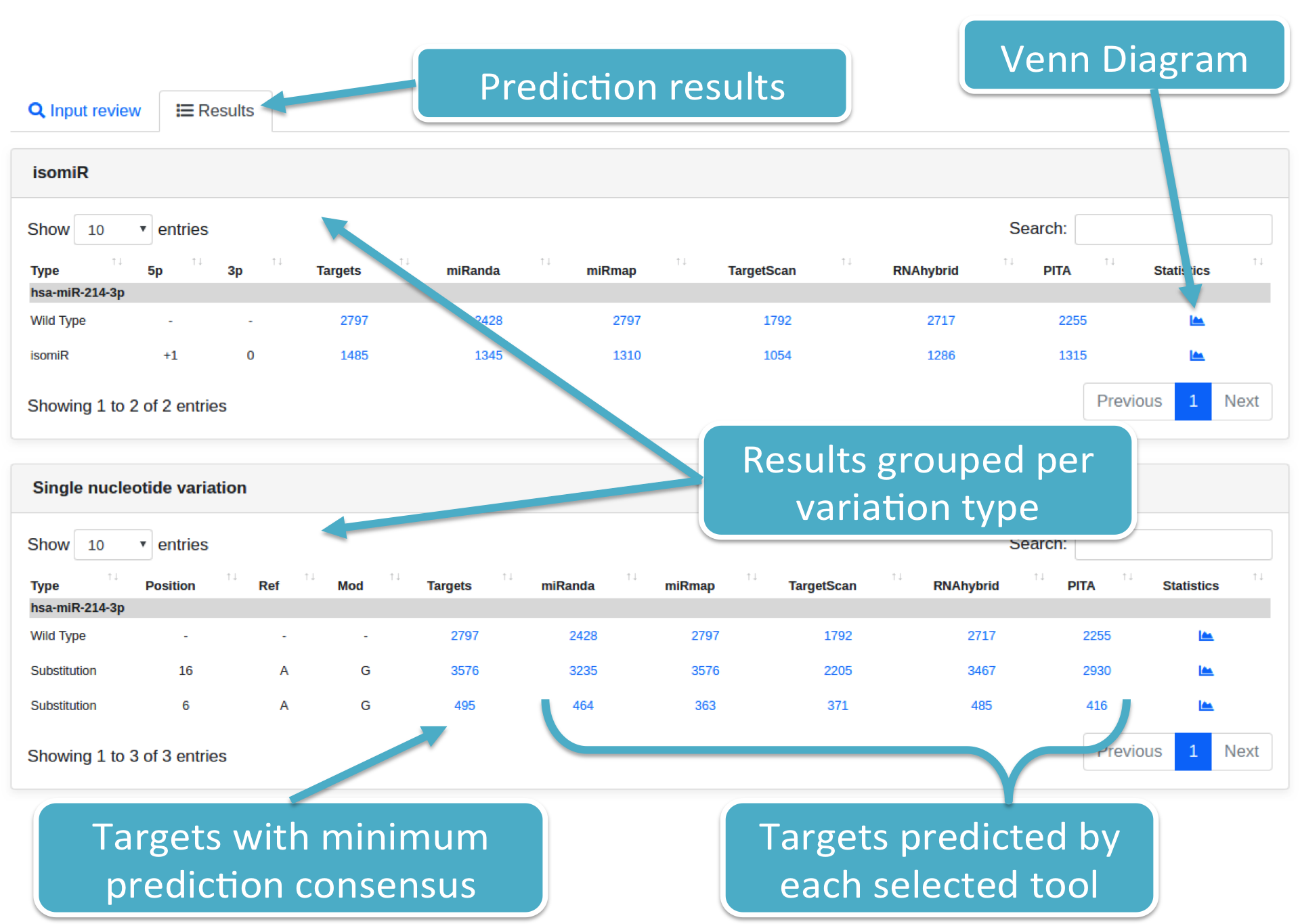
As shown above, isoTar adopts the same criterion for results visualization. In fact, prediction results are grouped per variation type, and results for each selected prediction tool are displayed. Each specified variation is associated with:
- Variation details, such as its type (i.e., Substitution or isomiR), position (for Substitution), and so on;
- The number of targets (named Targets), in which predicted targets are reported according to the specified minimum consensus, (0 in case there are none);
- The number of targets conclusion by each selected prediction tool (0 in case there are none);
- A Venn diagram reporting the targets intersection among all selected prediction tools.
Each number (colored in blue) represents the number of targets for a specific variation (i.e., Substitution or isomiR) and prediction tool/consensus. By clicking on it, the system will show you a modal box (figure below) reporting the targets list supplied with additional information. Results can be downloaded in different data formats, such as Excel, PDF, TSV, and so forth.
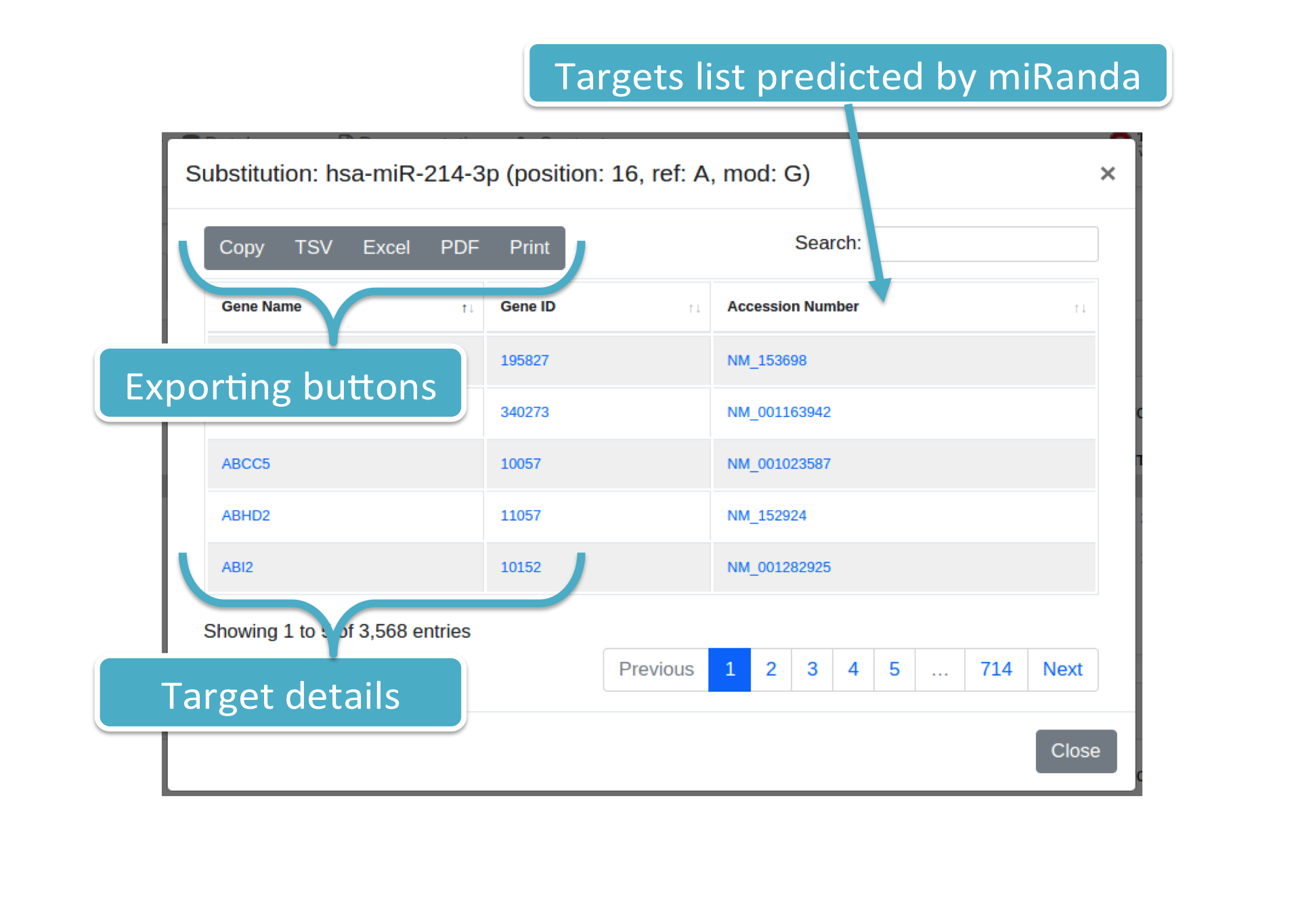
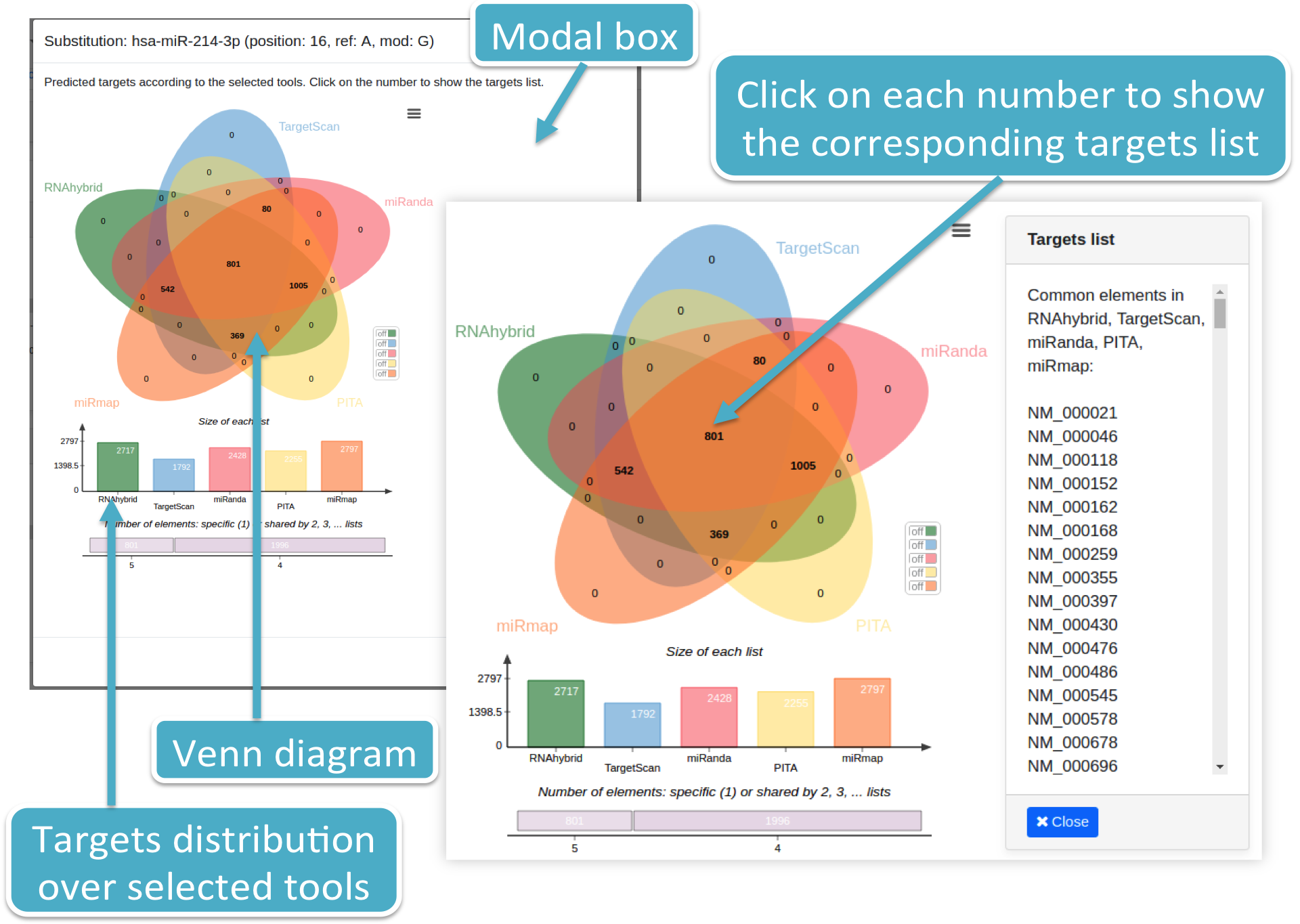
A Venn diagram is also provided, reporting the targets intersection among selected prediction tools, together with the histogram representing the targets distribution among such tools.
After the conclusion of the analysis (based on predicted targets), results are grouped according to each selected resource
(i.e., GO term, KEGG, Reactome), and details are displayed by variation type (i.e., Substitution, isomiR).
For each resource, the Functional Analysis is performed considering wild type, variation, and wild type vs. variation (comparison). Depending on which resource is
selected, isoTar provides different details. Below, we report results for the three main resources here available.
GO term
The GO term-based Functional Analysis is performed for each pair wild type-variation, specified as input for the Prediction Analysis. Then, results are grouped per Ontology category:
- Biological Process (BP);
- Cellular Component (CC);
- Molecular Function (MF).
A typical analysis provides information according to:
-
Wild Type
A GO term list for BP, CC, and MF, respectively, is supplied with additional details. Such GO terms are identified considering all the consensus predicted targets associated to the wild type miRNA; -
Variation
A GO term list for BP, CC, and MF, respectively, is supplied with additional details. Such GO terms are identified considering all the consensus predicted targets associated to the variation miRNA; -
Wild Type vs. Variation
-
A GO term list for each of the GO category (BP, CC, and MF) is supplied with additional details. GO terms are identified considering those consensus predicted targets that are associated with each of the following:
- Wild type only
- Variation only
- Wild type-variation intersection
-
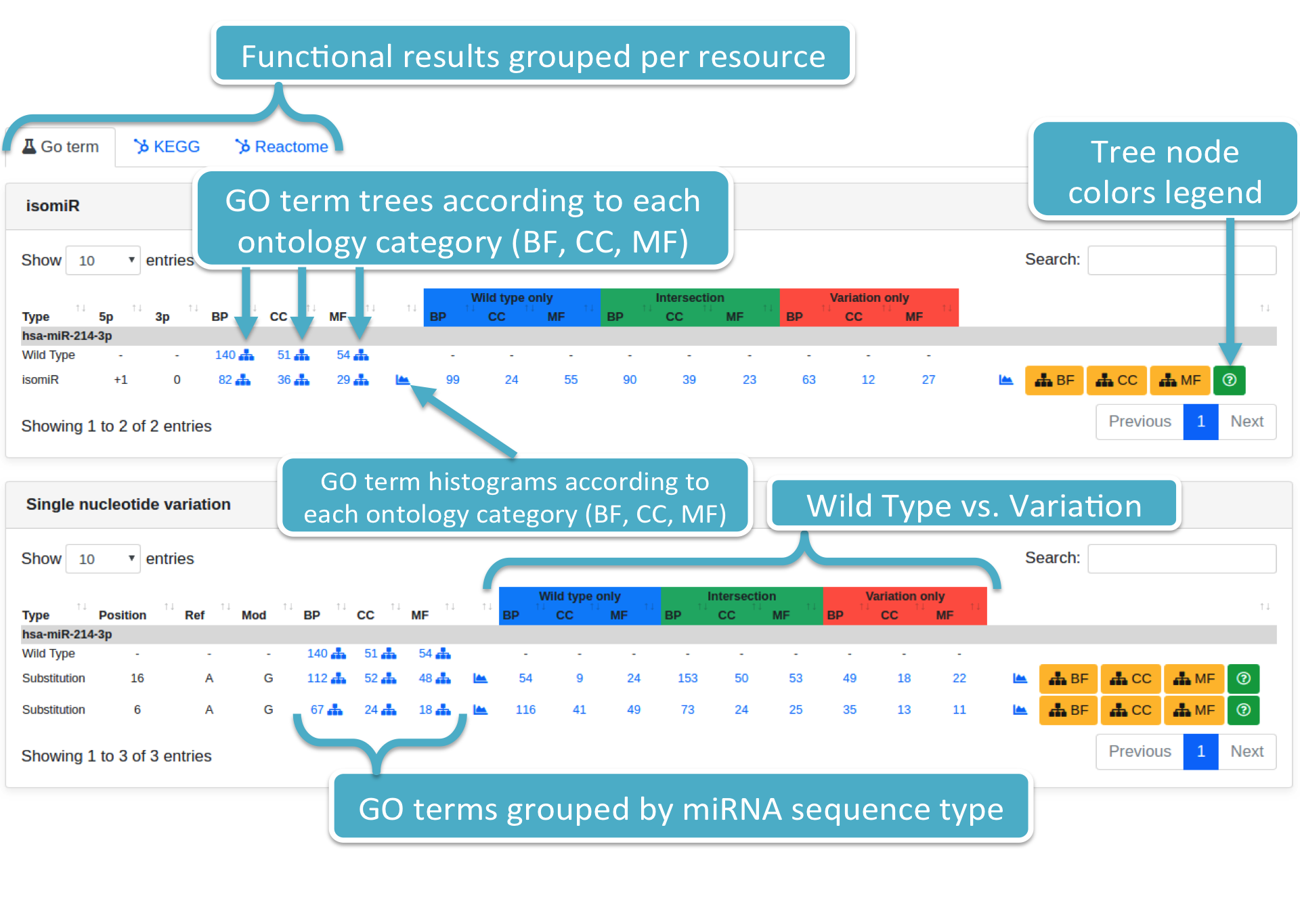
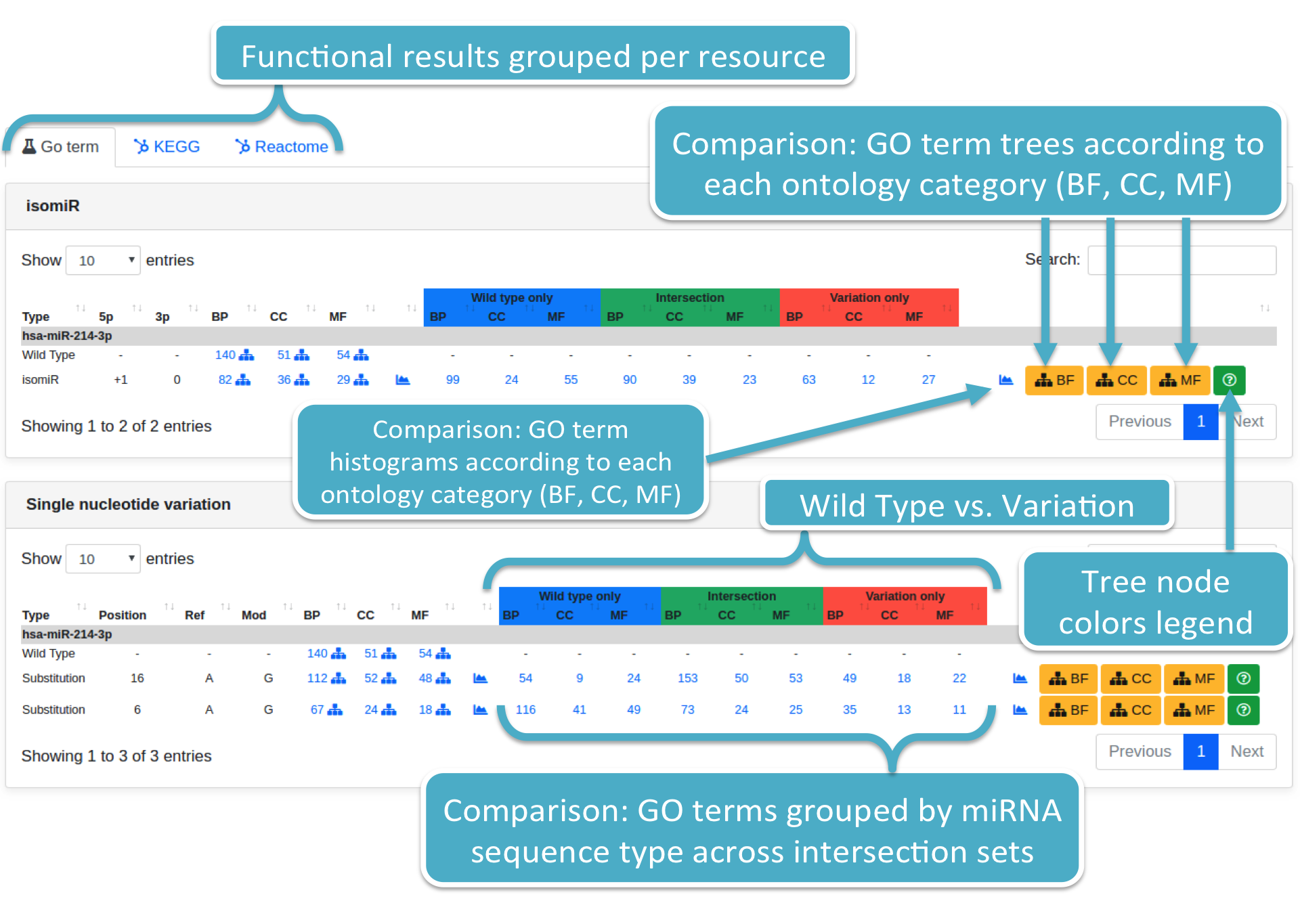
Results are displayed adopting the same format employed when displaying prediction results. Results are grouped per variation type and ontology category (i.e., BP, CC, MF). Each specified variation is associated with:
- Variation details, such as type (i.e., Substitution or isomiR), position (for Substitution), and so on;
- The number of identified GO terms (filtered per P-value type and threshold) for each ontology category, supplied with GO term trees and histograms;
- The number of GO terms (per ontology category) considering only those predicted targets associated to the wild type, variation, and their intersection, respectively. GO term trees and histograms are provided as well;
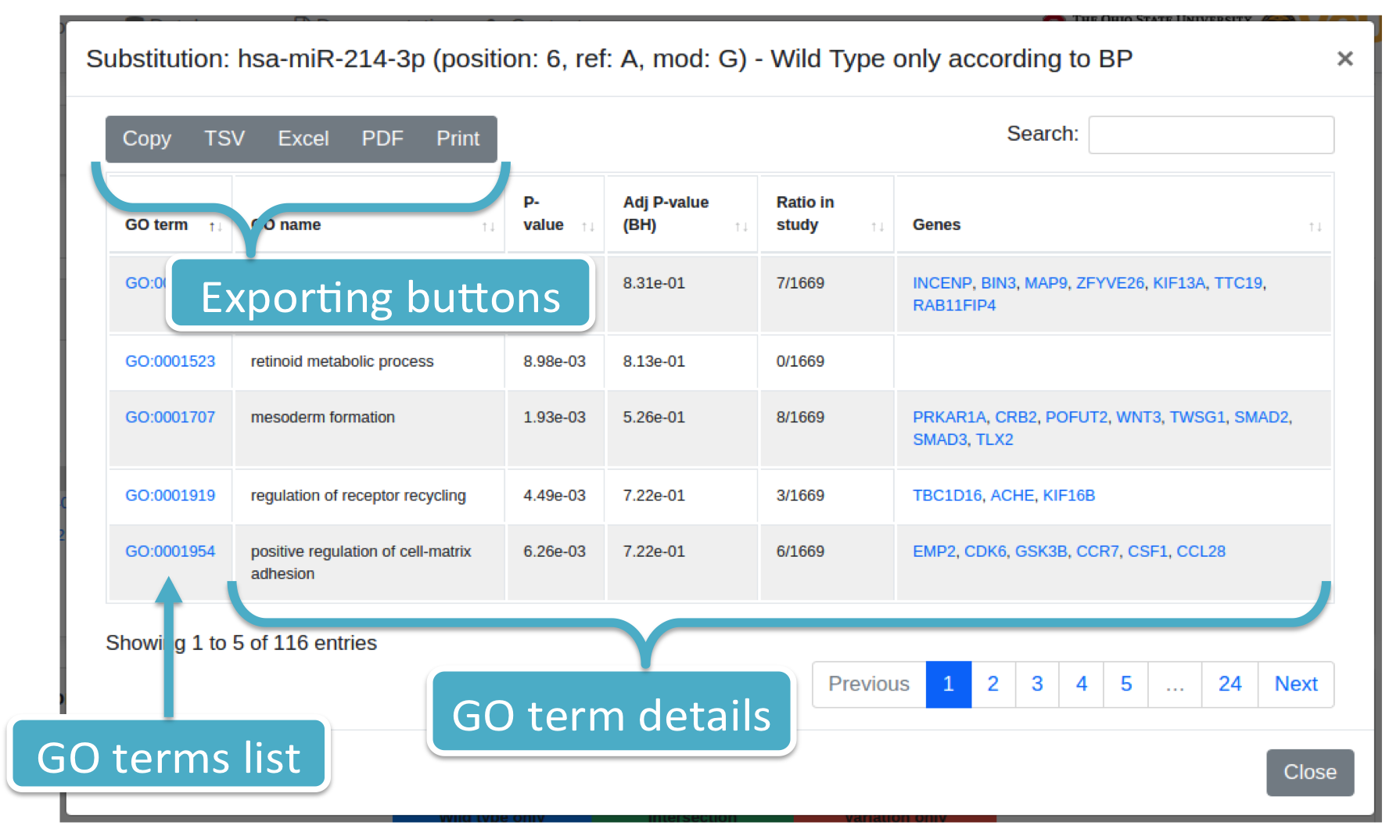
Each number (colored in blue) represents the number of GO terms for a specific wild type or variation (i.e., Substitution or isomiR) and ontology category (i.e., BP, CC, MF). By clicking on it, the system will show you a modal box (figure below) reporting the GO terms list supplied with additional information. Results can be downloaded in different data formats, such as Excel, PDF, TSV, and so forth.
Histograms are available for each ontology category. They report:
-
Comparison between wild type and variation. As an example, we report (figure below) a histogram associated with the wild type and the variation at
position 6 according to the ontology category CC (wild type in blue and variation in red);
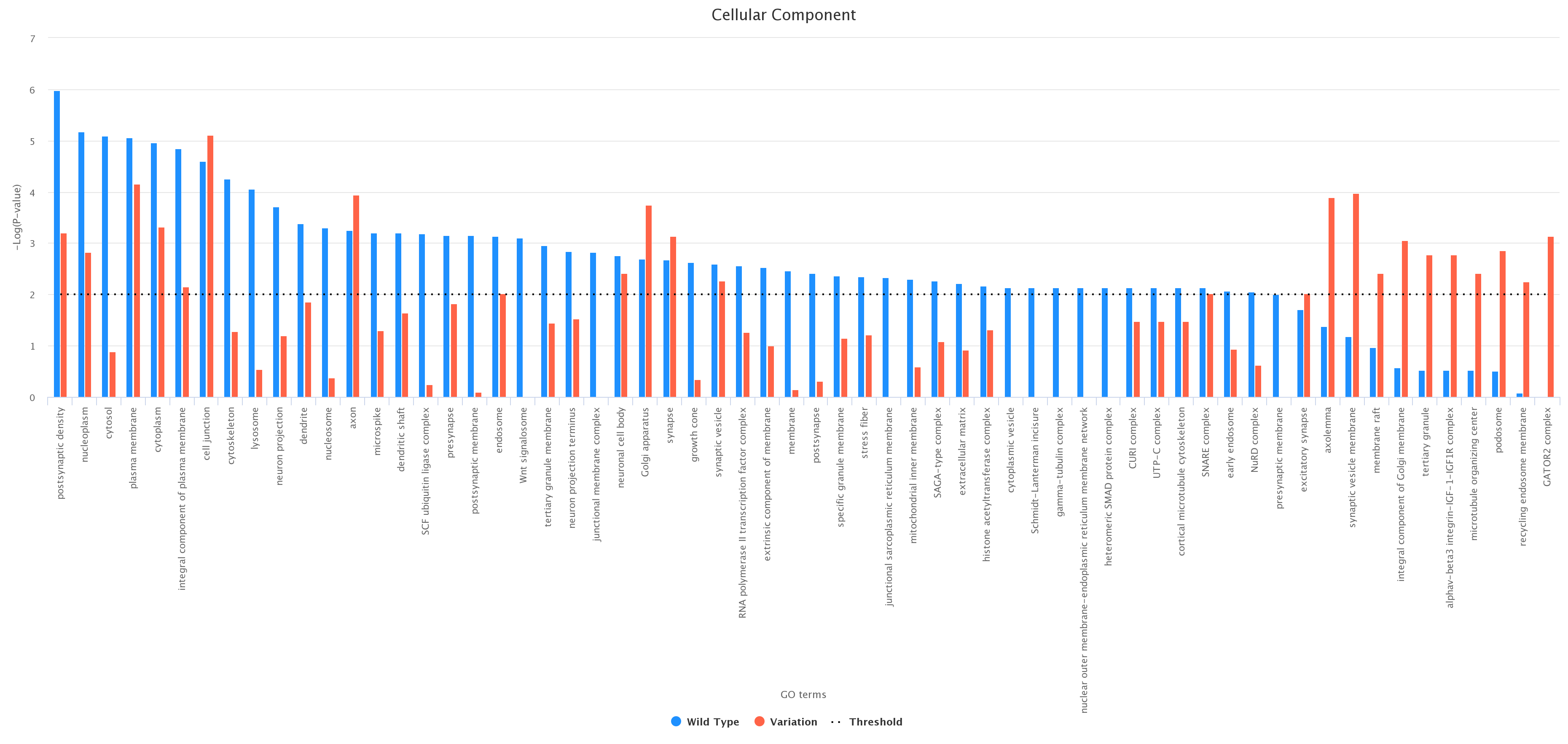
-
Comparison between wild type only, variation only, and their intersection. As an example, we report (figure below) a histogram associated with the comparison between the wild type and the variation (wild type only, variation only, and their intersection) at
position 6 according to the ontology category CC (wild type in blue, variation in red, and intersection in green);
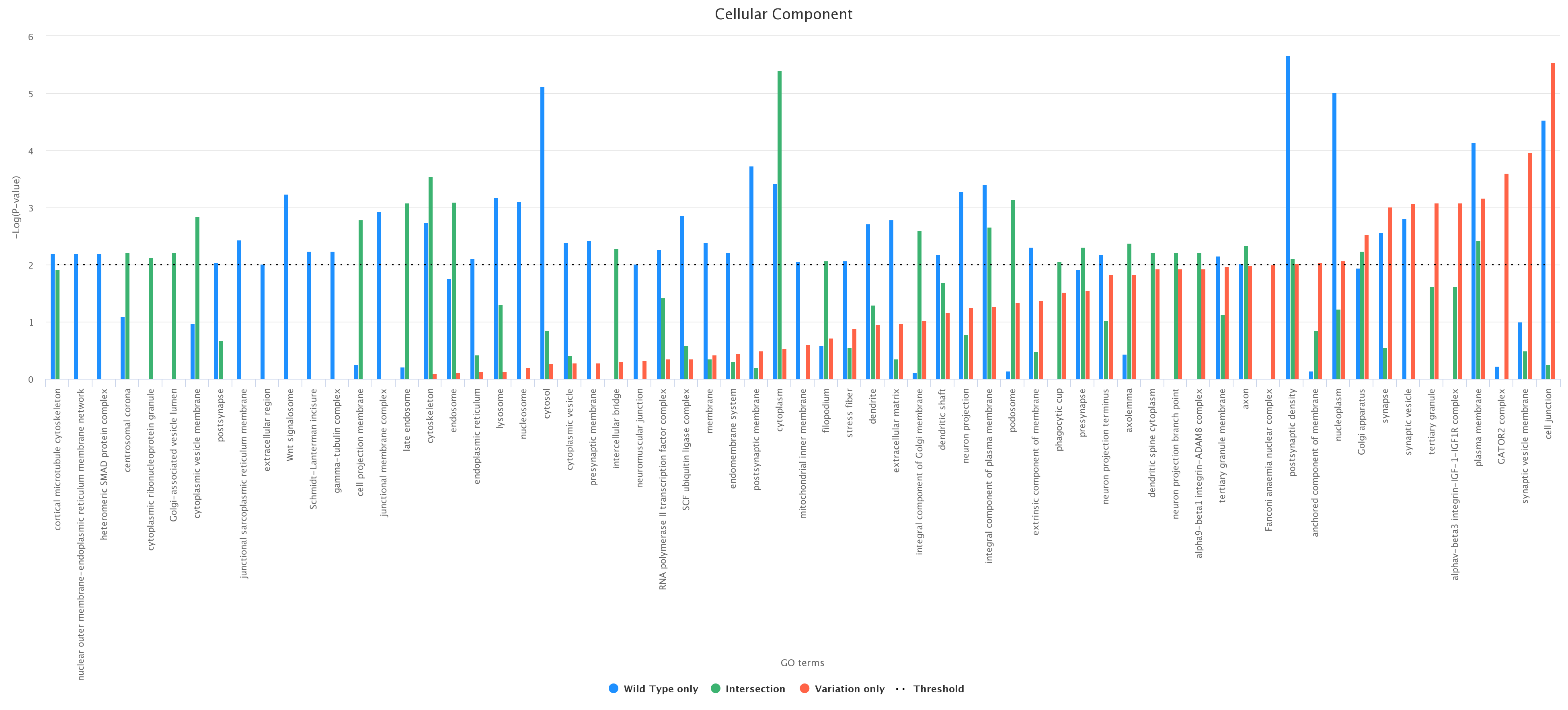
Histograms can be downloaded in different formats, such as PDF, PNG, JPEG, and SVG.
GO term trees are available for each ontology type. They are associated with:
-
Each wild type and their variation(s). As an example, we report (figure below) two GO term trees associated with the wild type and the variation at
position 6, respectively, according to the ontology type CC (in green, the GO term nodes for which the P-value is also reported);
Wild Type 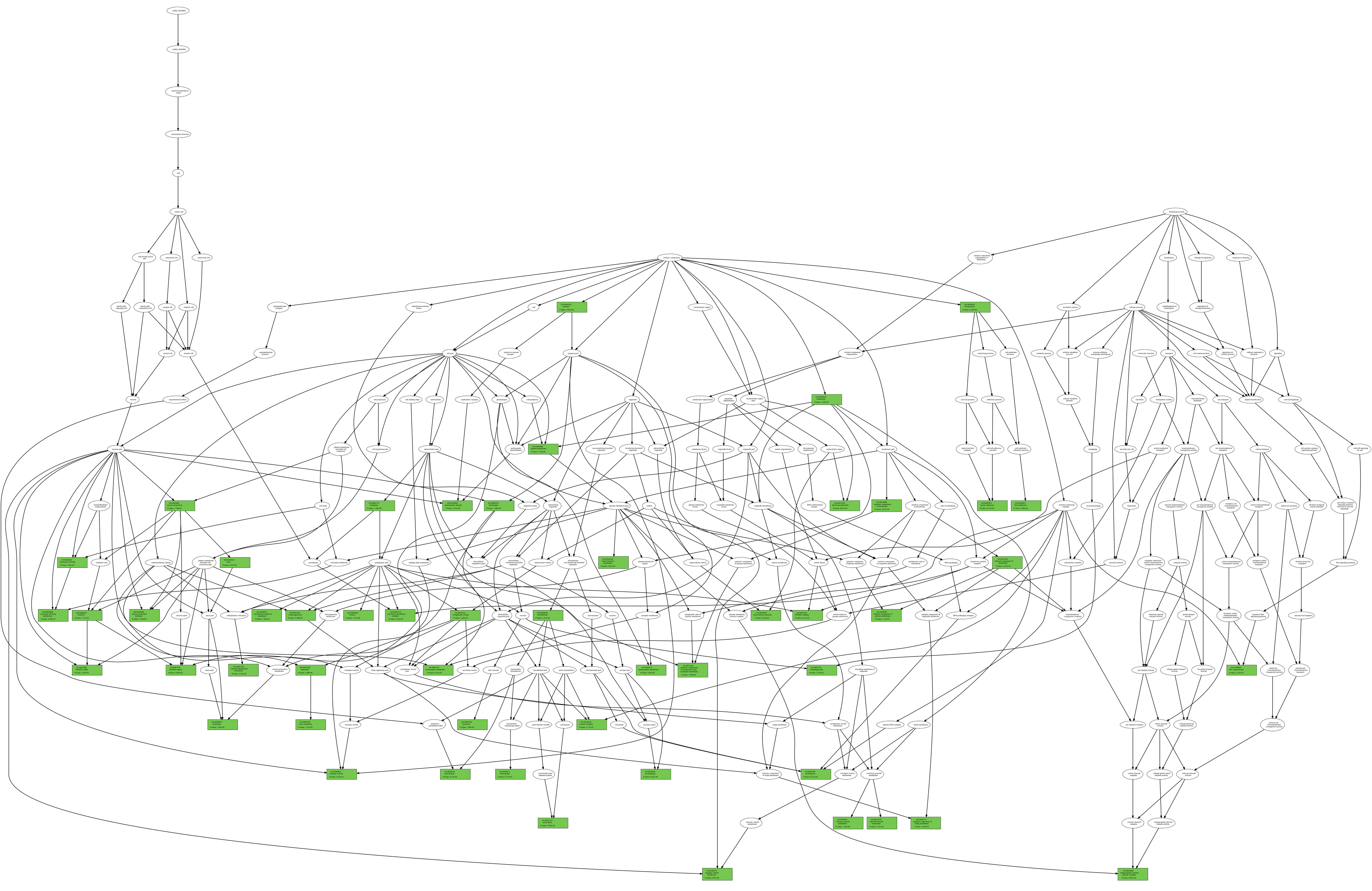
Variation 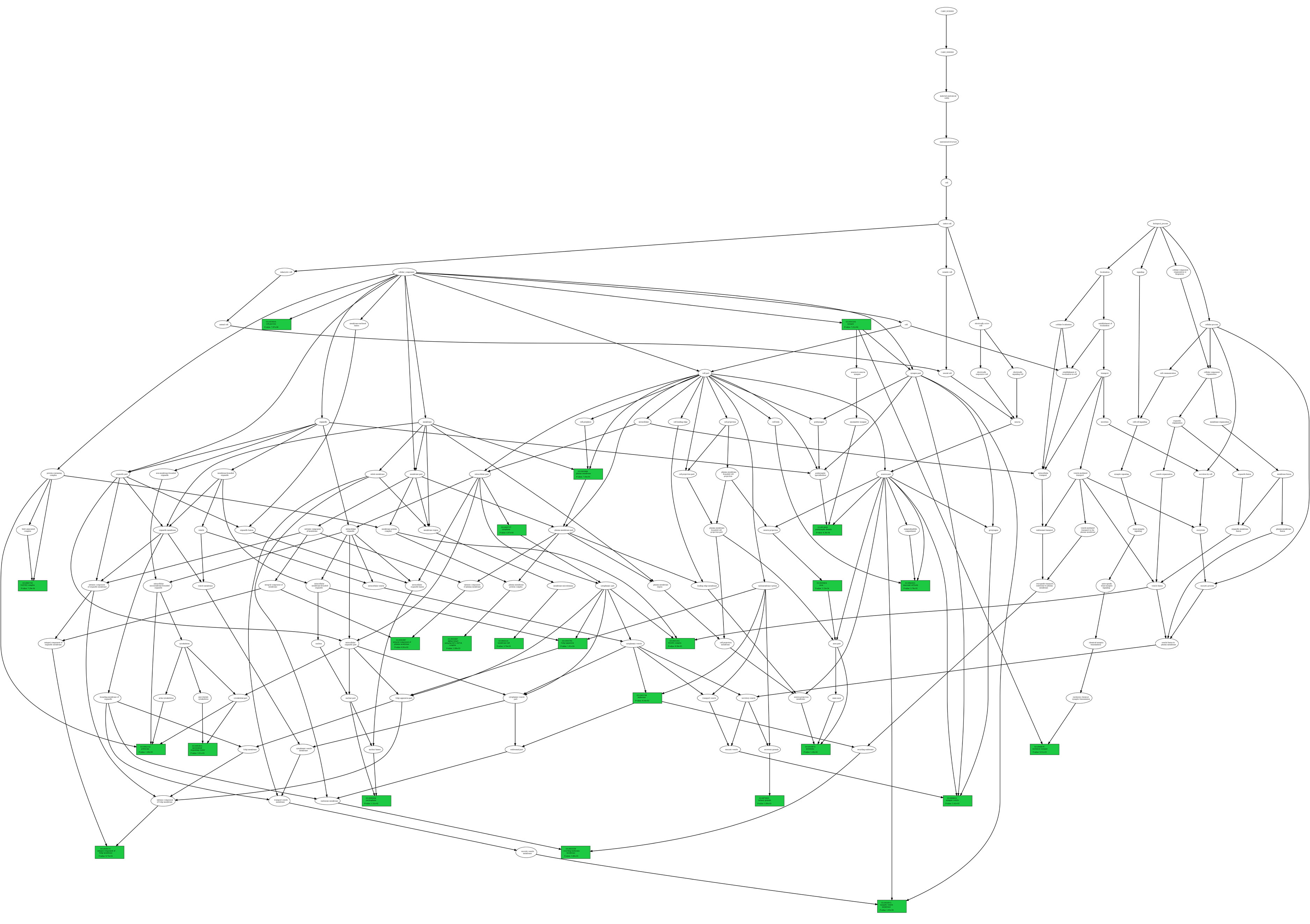
-
Comparison between wild type only, variation only, and their intersection. As an example, we report (figure below) the GO term tree associated with the comparison
between the wild type and the variation (wild type only, variation only, and their intersection) at position 6 according to the ontology type CC;
Wild Type vs. Variation 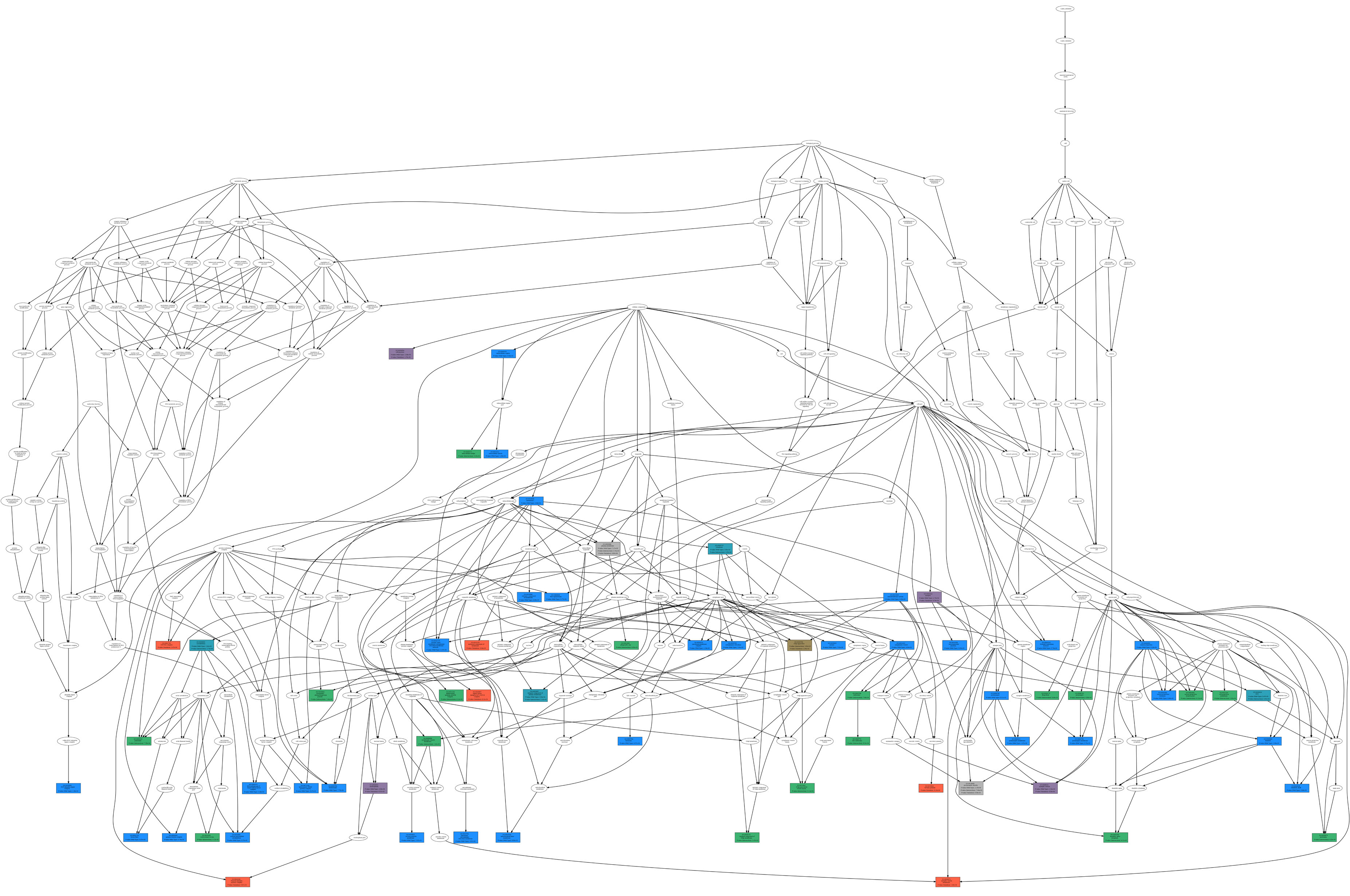
Colors are used to highlight GO term nodes, according to all possible combinations among all possible intersection subsets. A table reporting the adopted colors is provided as well (figure below).
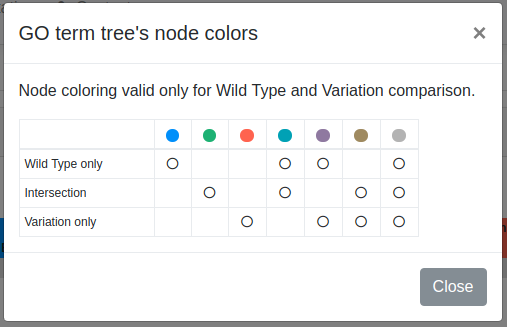
GO term trees can be downloaded in PDF format.
KEGG & Reactome
The Pathway-based Functional Analysis is performed for each pair wild type-variation, specified as input for the Prediction Analysis. A typical analysis provides information according to:-
Wild Type
A pathways list is supplied with additional details. Pathways are identified considering all consensus predicted targets associated to the wild type; -
Variation
A pathways list is supplied with additional details. Pathways are identified considering all consensus predicted targets associated to the variation; -
Wild Type vs. Variation
-
A pathways list is supplied with additional details. Pathways are identified considering those consensus predicted targets that are respectively associated only to:
- Wild type
- Variation
- Wild type-variation intersection
-
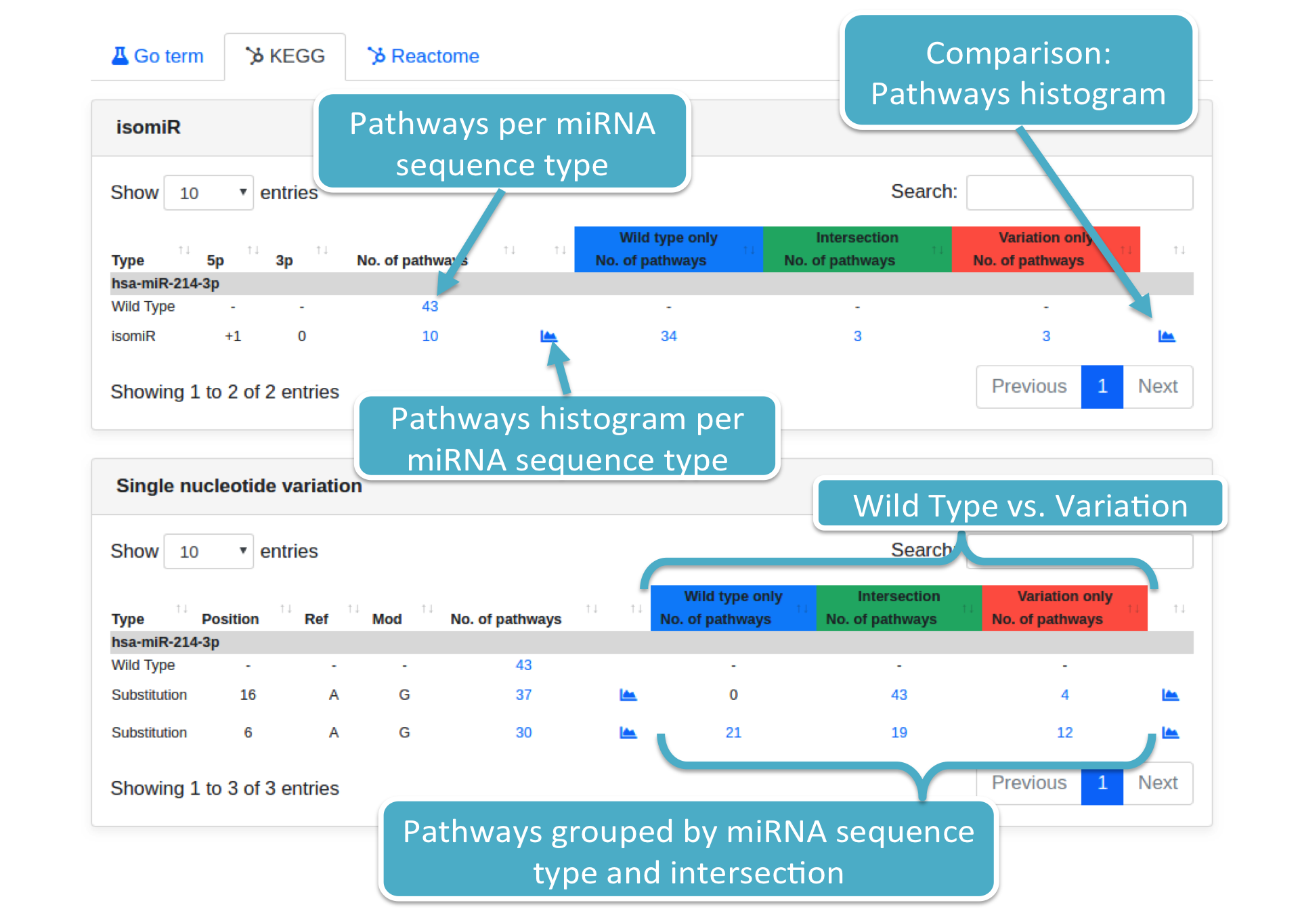
Results are displayed adopting the same format employed when displaying prediction results (grouped per variation type). Each specified variation is associated with:
- Variation details, such as type (i.e., Substitution or isomiR), position (Substitution only), and so on;
- The number of identified pathways (filtered by P-value type and threshold), supplied with histograms;
- The number of pathways considering only those predicted targets associated with the wild type, variation, and their intersection, respectively. Histograms are provided as well;
Each number (colored in blue) represents the number of pathways for a specific variation (i.e., Substitution or isomiR). By clicking on it, the system will show you a modal box (figure below) reporting the pathways list supplied with additional information. Results can be downloaded in different data formats, such as Excel, PDF, TSV, and so forth.
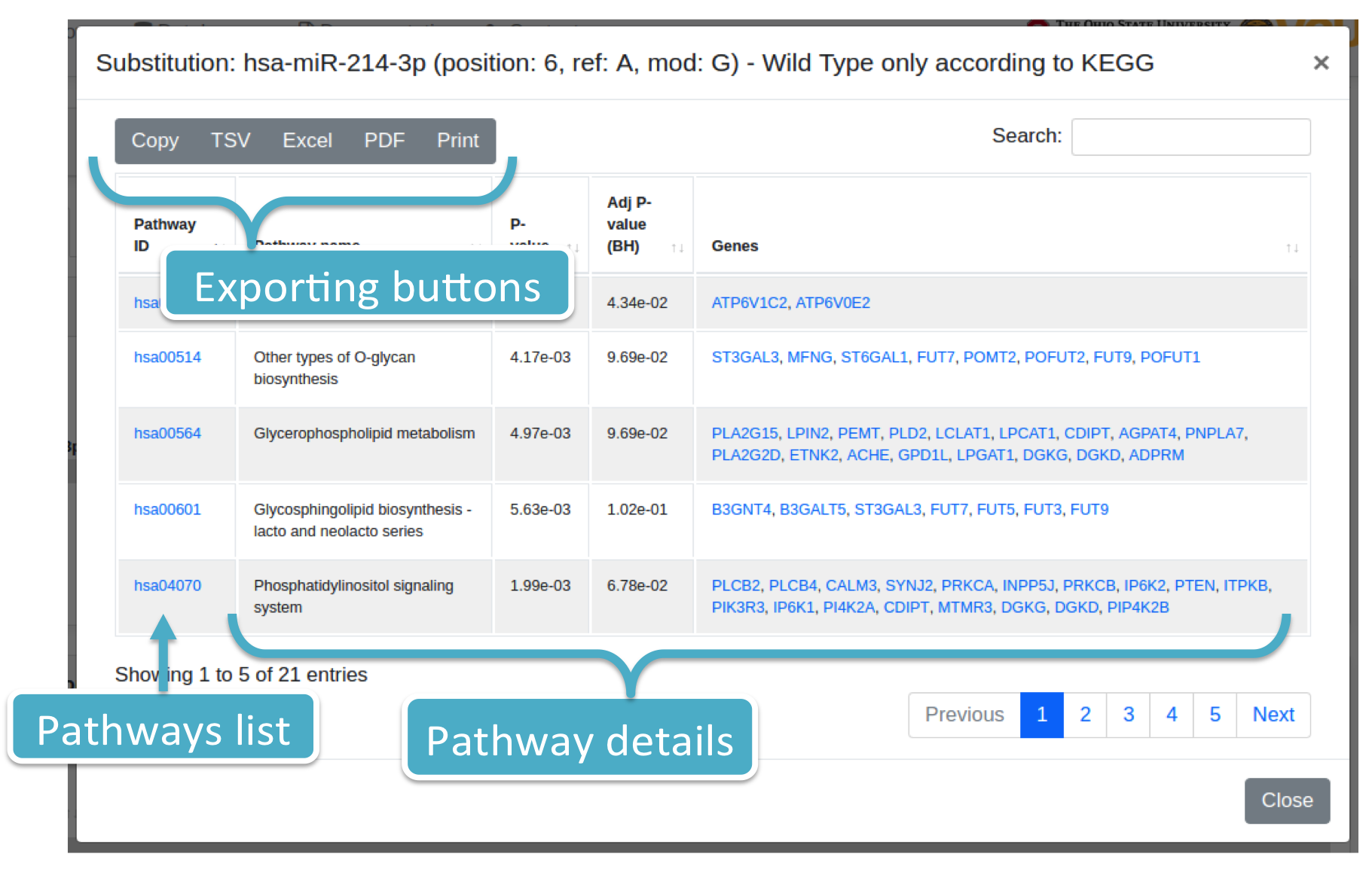
Histograms report:
-
Comparison between wild type and variation. As an example, we report (figure below) a histogram associated with the wild type and the variation at
position 6 according to KEGG (wild type in blue and variation in red);
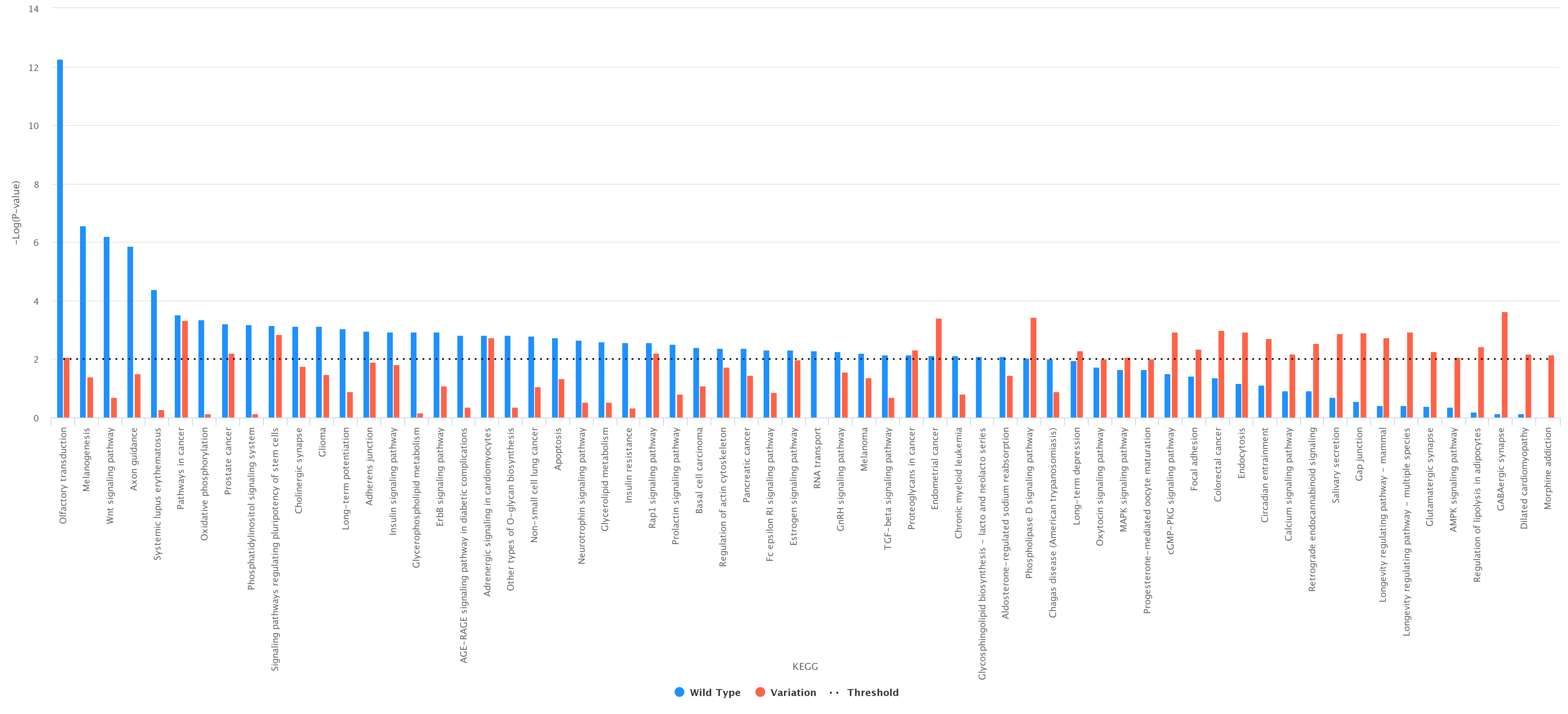
-
Comparison between wild type only, variation only, and their intersection. As an example, we report (figure below) a histogram associated with the comparison between the wild type and the variation (wild type only, variation only, and their intersection) at
position 6 according to KEGG (wild type in blue, variation in red, and intersection in green);
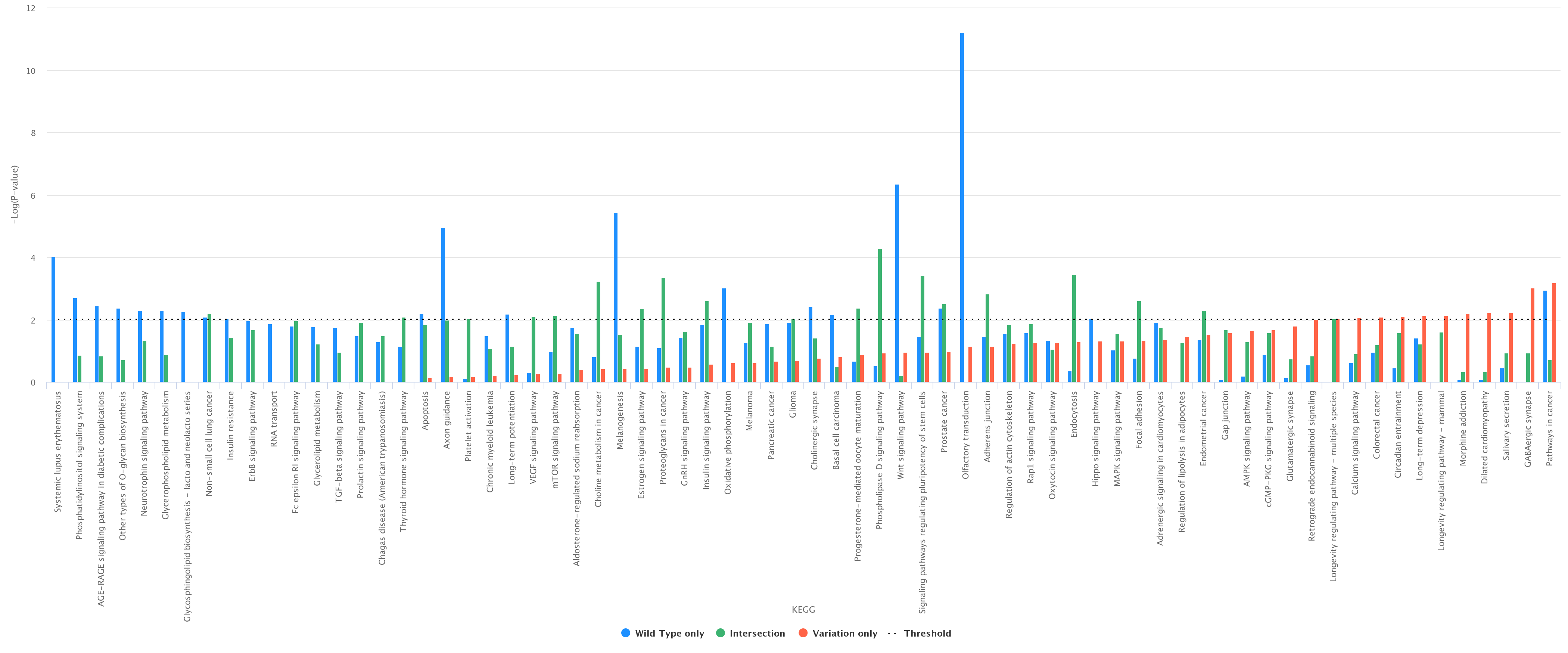
Histograms can be downloaded in different formats, such as PDF, PNG, JPEG, and SVG.
What is described above is valid for Reactome results al well.
When performing Functional Analysis without leveraging on consensus predicted targets, one or two user-specified lists of targets may be provided. In this section,
we describe results for two lists of targets. For clarity, we named the first list wild_type (List_1) and the second one variation
(List_2). After the conclusion of the analysis, results are grouped according to each selected resource (i.e., GO term, KEGG, Reactome).
For each resource, the Functional Analysis is performed considering List_1, List_2, and List_1 vs. List_2 (comparison). Depending on which resource is
selected, isoTar provides different details. Below, we report results for the three main resources here available.
GO term
The GO term-based Functional Analysis is performed for List_1 and List_2. Then, the results are grouped per Ontology category:
- Biological Process (BP);
- Cellular Component (CC);
- Molecular Function (MF).
A typical analysis provides information according to:
-
List_1
A GO terms list for BP, CC, and MF, respectively, is supplied with additional details. Such GO terms are identified considering all List_1 targets; -
List_2
A GO terms list for BP, CC, and MF, respectively, is supplied with additional details. Such GO terms are identified considering all List_2 targets; -
List_1 vs. List_2
-
A GO term list for each of the GO category (BP, CC, MF) is supplied with additional details. GO terms are identified considering those consensus predicted targets that are associated with each of the following:
- List_1
- List_2
- List_1-List_2 intersection
-
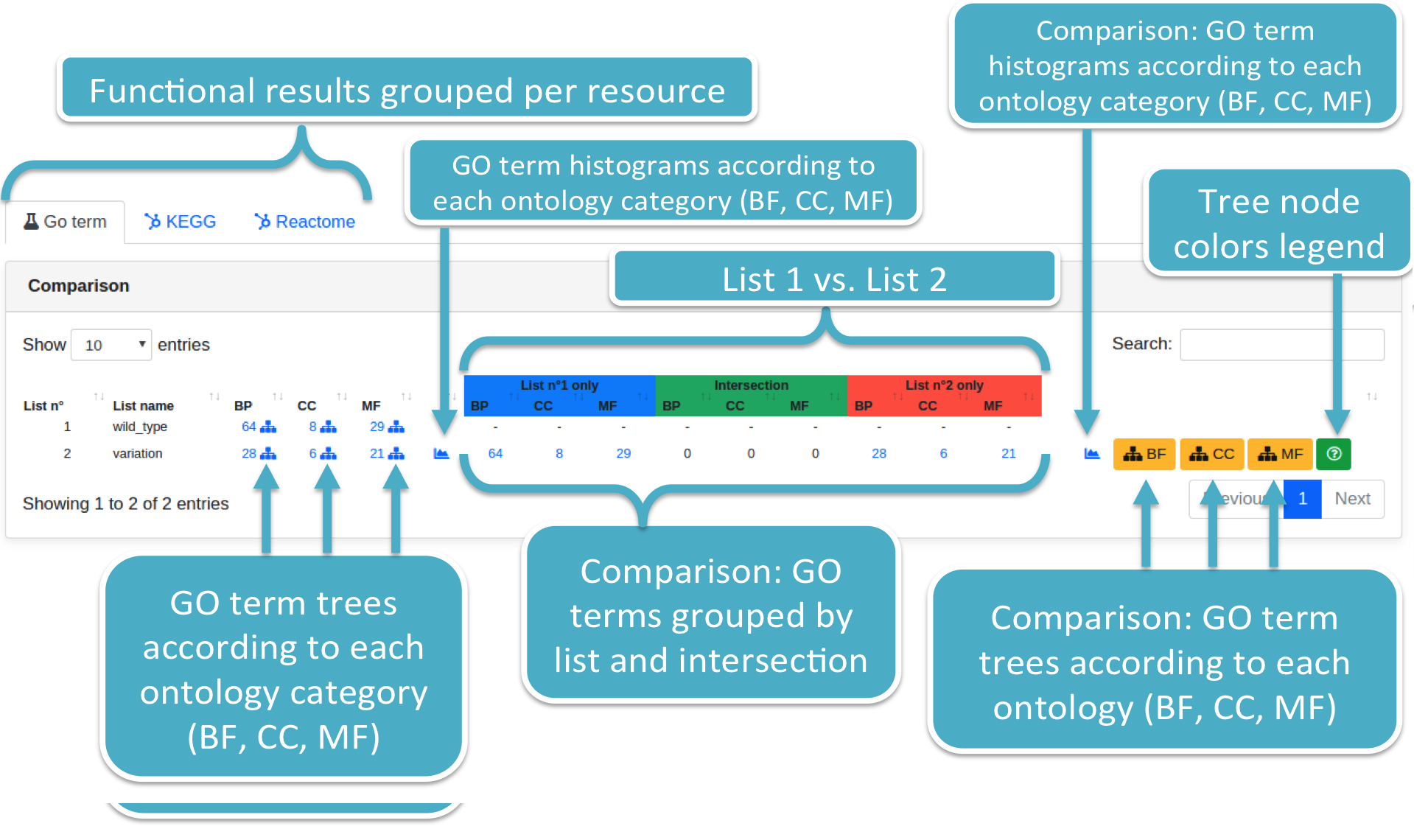
Results are displayed adopting the same format employed when displaying prediction results. Results are grouped per ontology category (i.e., BP, CC, MF). List_2 is associated with:
- The number of identified GO terms (filtered per P-value type and threshold) for each ontology category, supplied with GO term trees and histograms;
- The number of GO terms (per ontology category) considering only those targets associated to the List_1, List_2, and their intersection, respectively. GO term trees and histograms are provided as well.
Each number (colored in blue) represents the number of GO terms for List_1 or List_2 and ontology category (i.e., BP, CC, MF). By clicking on it, the system will show you a modal box (figure below) reporting the GO terms list supplied with additional information. Results can be downloaded in different data formats, such as Excel, PDF, TSV, and so forth.
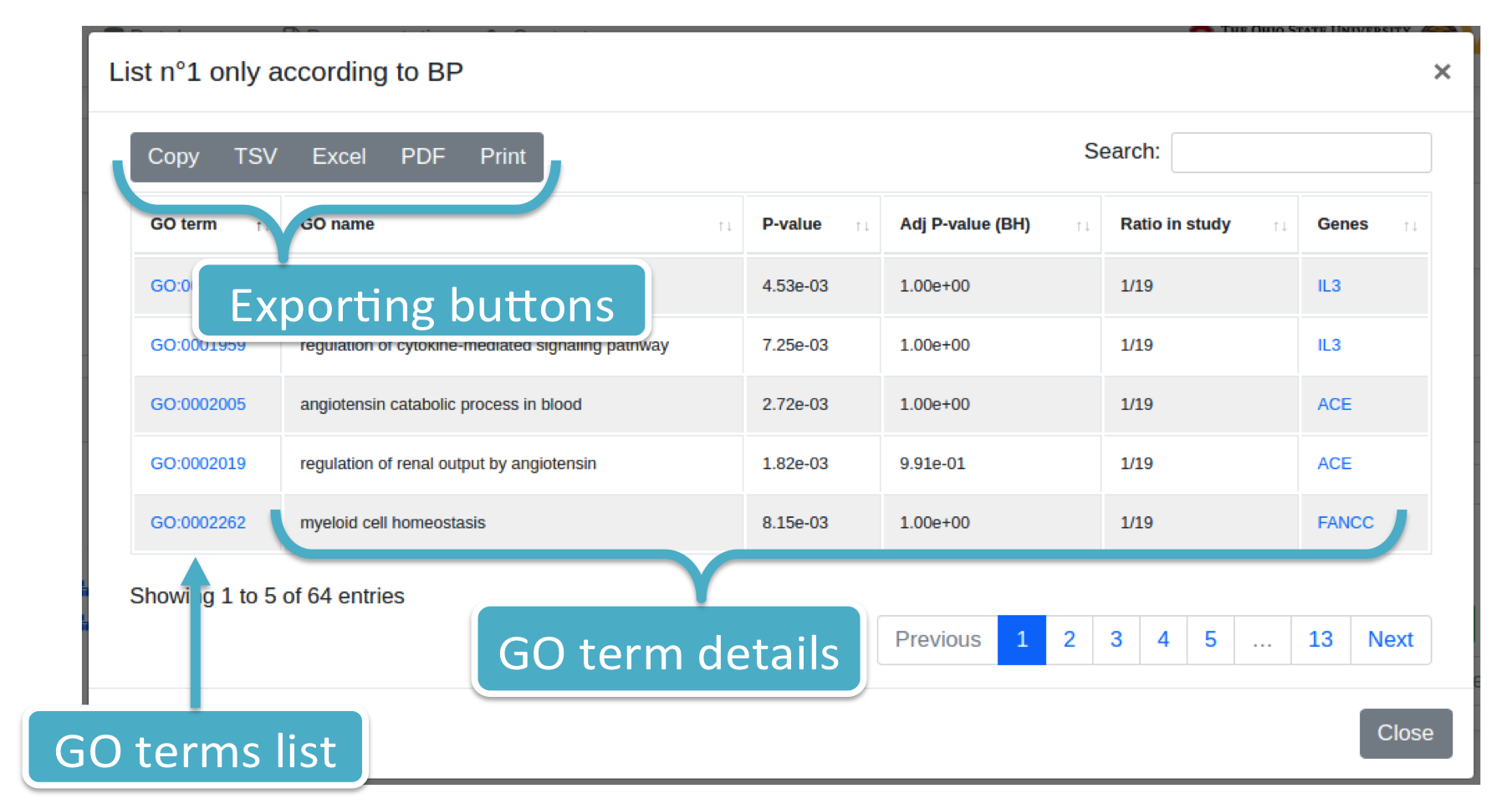
Histograms are available for each ontology category. They report:
-
Comparison between List_1 and List_2. As an example, we report (figure below) a histogram associated with List_1 and List_2 according to
the ontology category CC (wild type in blue and variation in red);
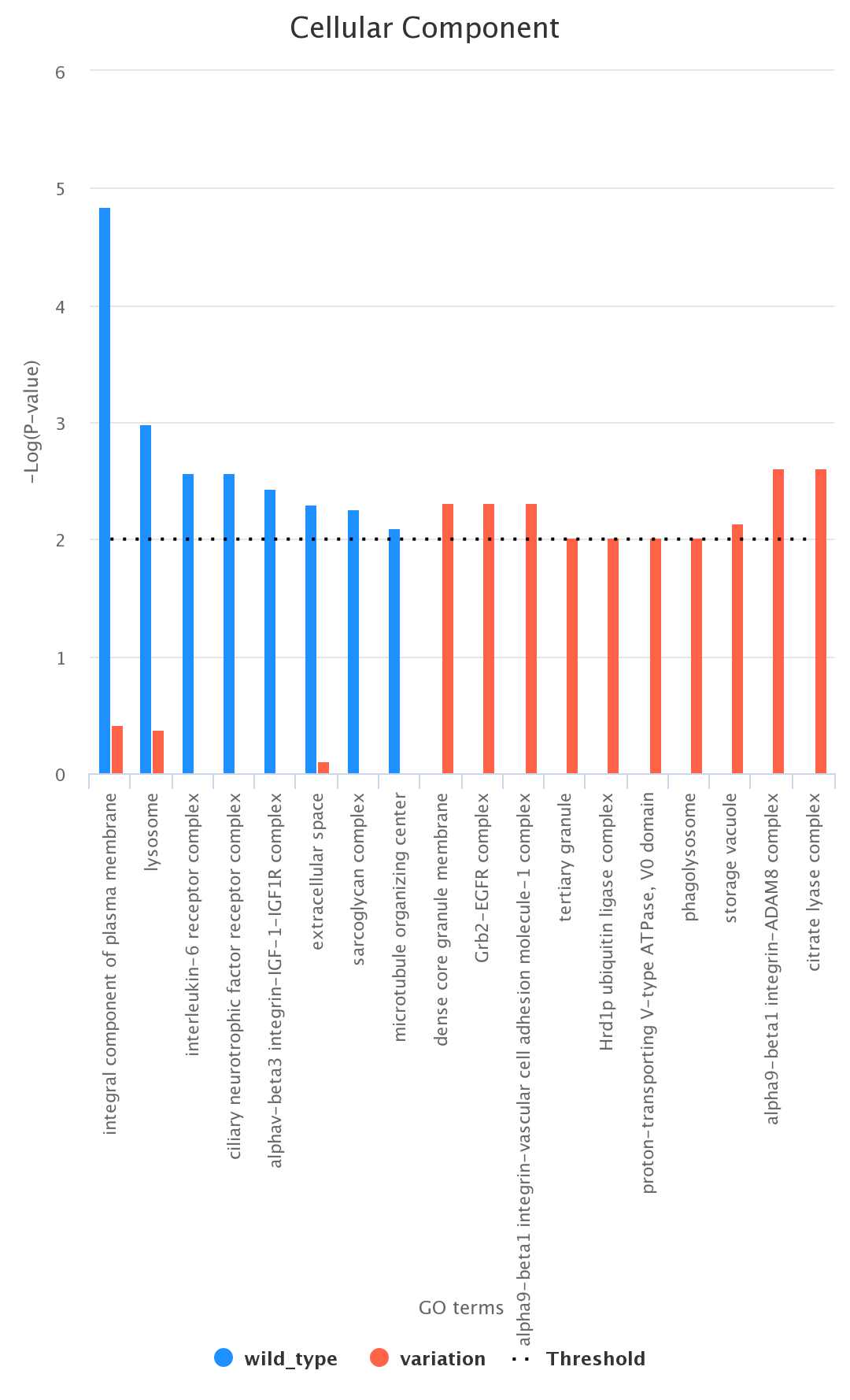
-
Comparison between List_1 only, List_2 only, and their intersection. As an example, we report (figure below) a histogram associated with the
comparison between the List_1 and the List_2 (wild type only, variation only, and their intersection) according to the ontology category CC
(wild type in blue, variation in red, and intersection in green);
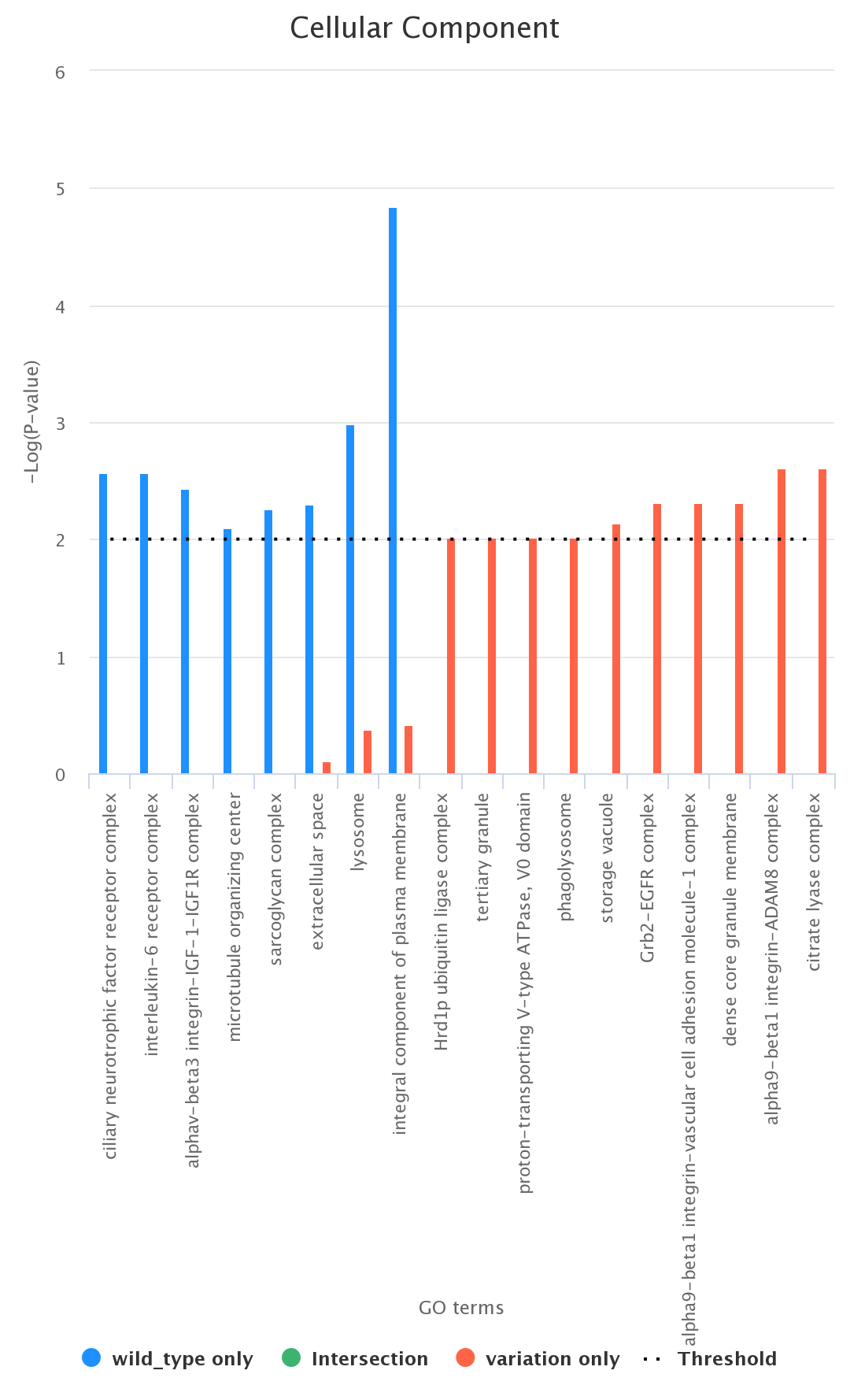
Histograms can be downloaded in different formats, such as PDF, PNG, JPEG, and SVG.
GO term trees are available for each ontology category. They are associated with:
-
List_1 and List_2. As an example, we report (figure below) two GO term trees associated with List_1 and List_2, respectively, according to
the ontology category CC (in green, the GO term nodes for which the P-value is also reported);
List_1 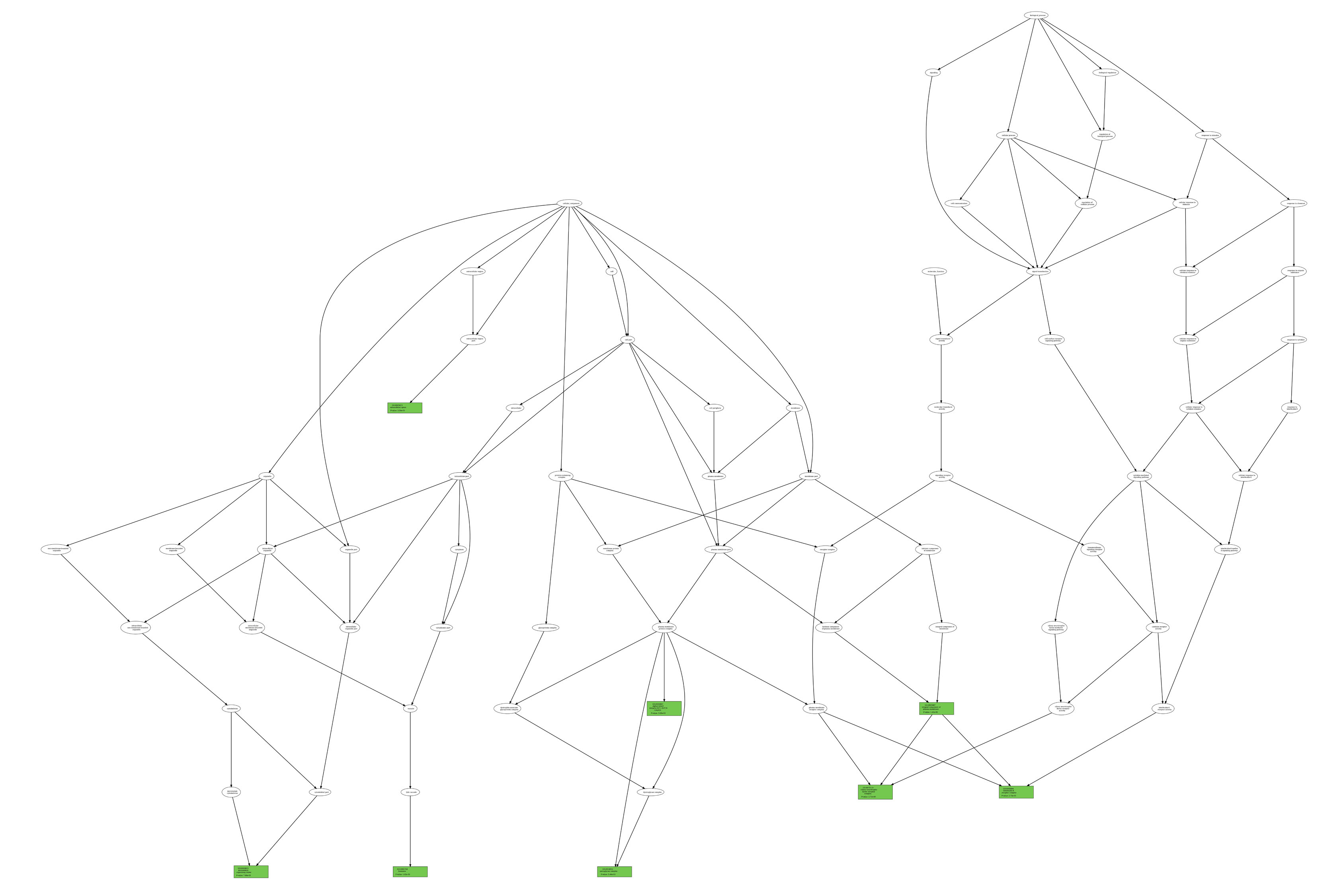
List_2 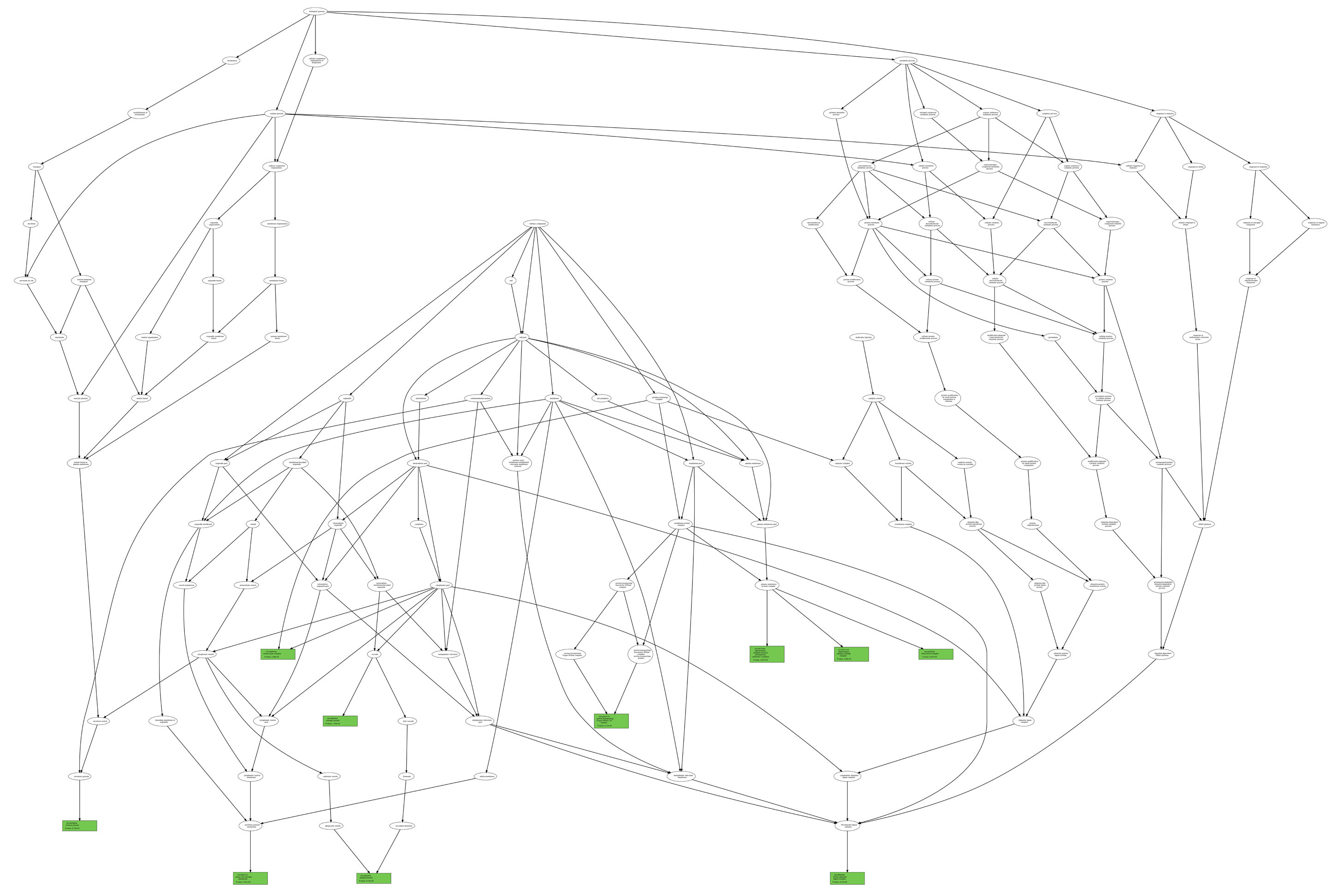
-
Comparison between List_1 only, List_1 only, and their intersection. As an example, we report (figure below) the GO term tree associated with the
comparison between List_1 and List_2 (List_1 only, List_2 only, and their intersection) according to the ontology category CC;
List_1 vs. List_2 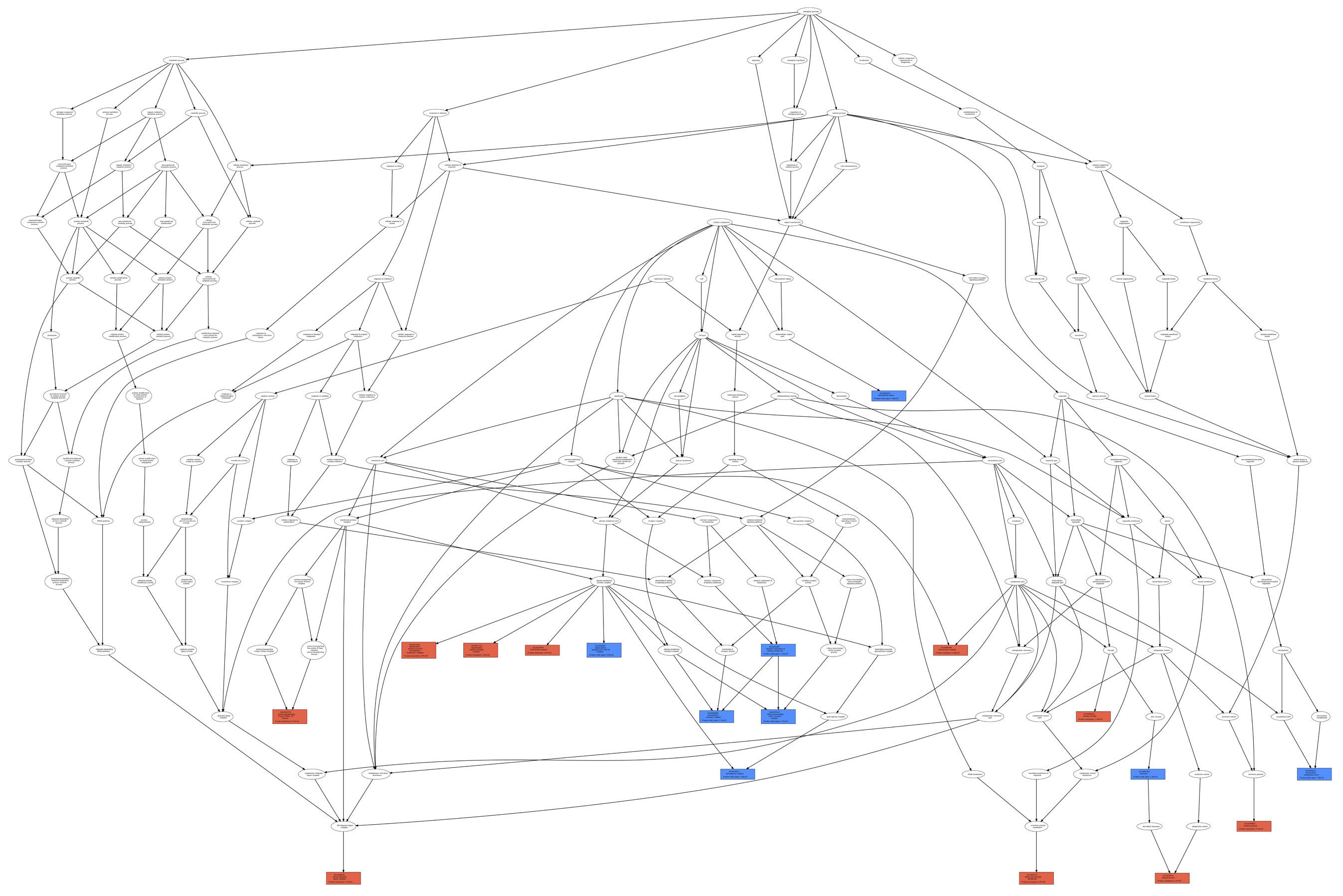
Colors are used to highlight GO term nodes, according to all possible combinations among all possible intersection subsets. A table reporting the adopted colors is provided as well (figure below).
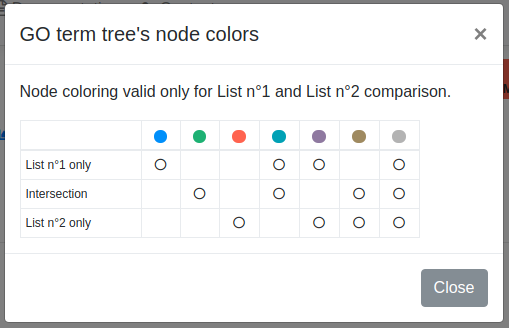
GO term trees can be downloaded in PDF format.
KEGG & Reactome
The Pathway-based Functional Analysis is performed considering the pair List_1-List_2. A typical analysis provides information according to:
-
List_1
A pathways list is supplied with additional details. Pathways are identified considering all List_1 targets; -
List_2
A pathways list is supplied with additional details. Pathways are identified considering all List_2 targets; -
List_1 vs. List_2
-
A pathways list is supplied with additional details. Pathways are identified considering those consensus predicted targets that are respectively associated only to:
- List_1
- List_2
- List_1-List_2 intersection
-
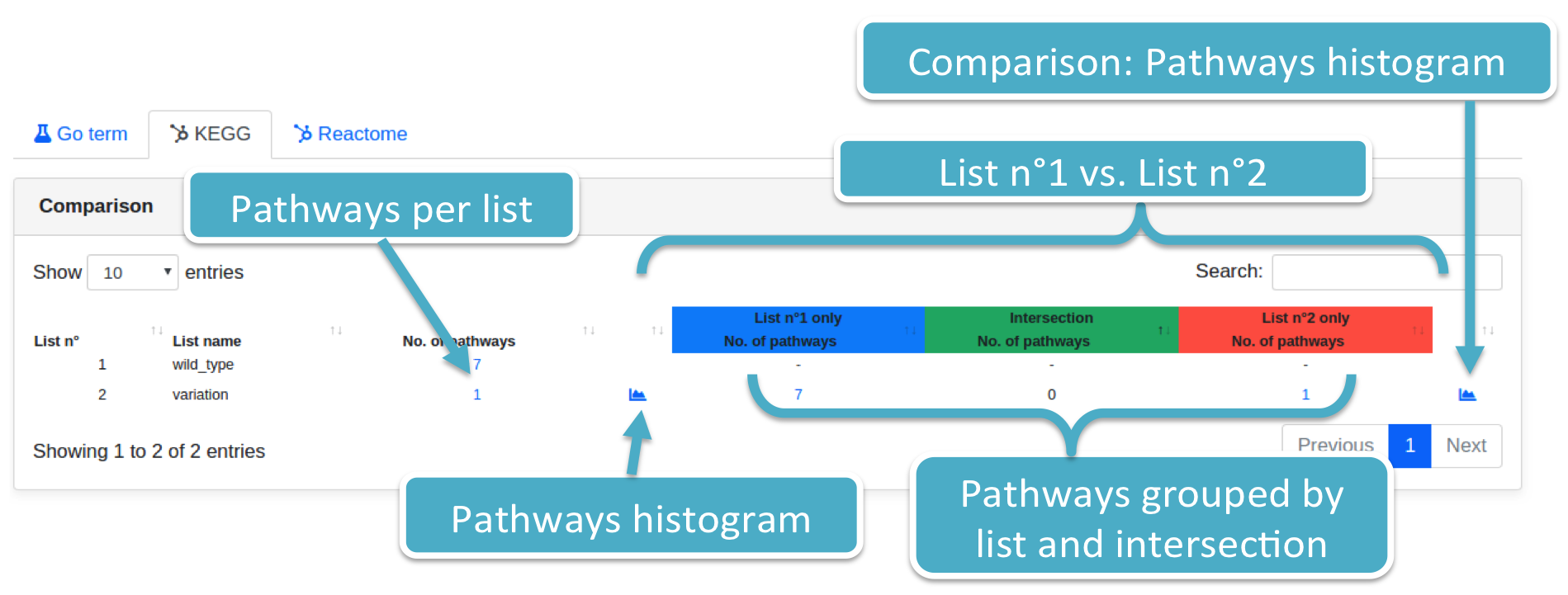
List_2 is associated with:
- The number of identified pathways (filtered per P-value type and threshold), supplied with histograms;
- The number of pathways considering only those targets associated with List_1, List_2, and their intersection. Histograms are provided as well;
Each number (colored in blue) represents the number of pathways. By clicking on it, the system will show you a modal box (figure below) reporting the pathways list supplied with additional information. Results can be downloaded in different data formats, such as Excel, PDF, TSV, and so forth.
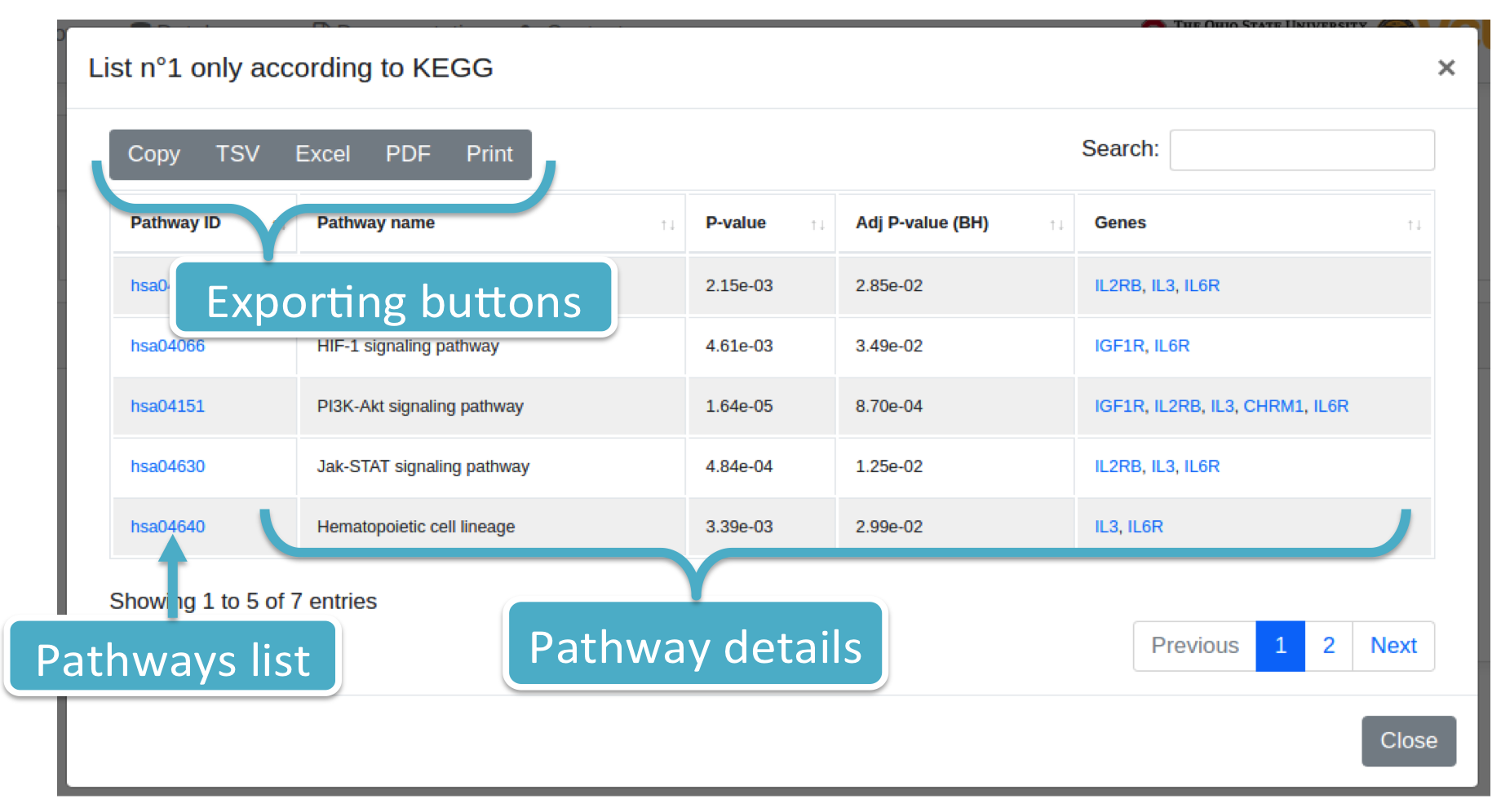
Histograms report:
-
Comparison between List_1 and List_2. As an example, we report (figure below) a histogram associated with List_1 and List_2 according to KEGG
(List_1 in blue and List_2 in red);
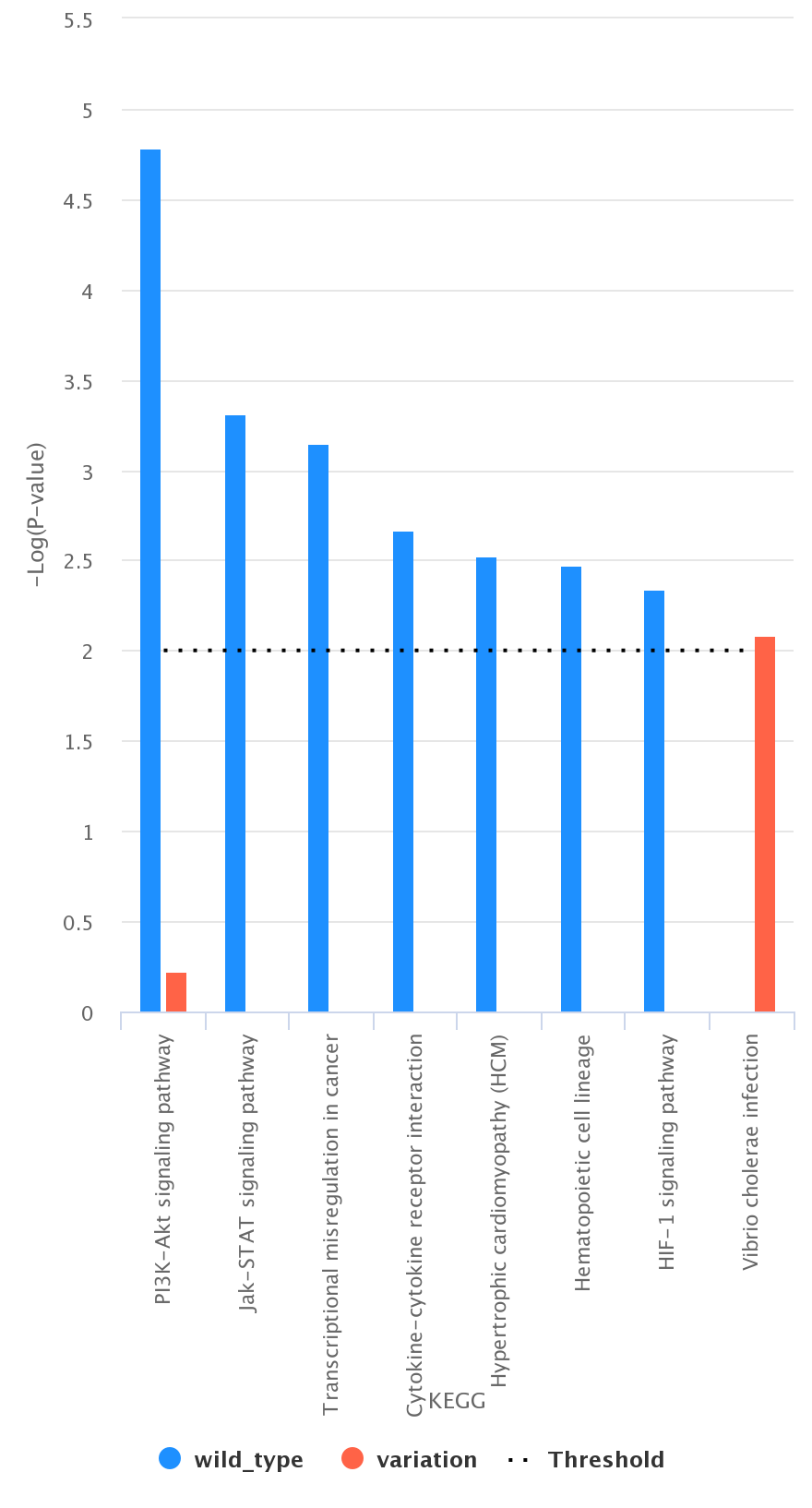
-
Comparison between List_1 only, List_2 only, and their intersection. As an example, we report (figure below) a histogram associated with the comparison
between List_1 and List_2 (List_1 only, List_2 only, and their intersection) according to KEGG (List_1 in blue, List_2 in red, and intersection in green);
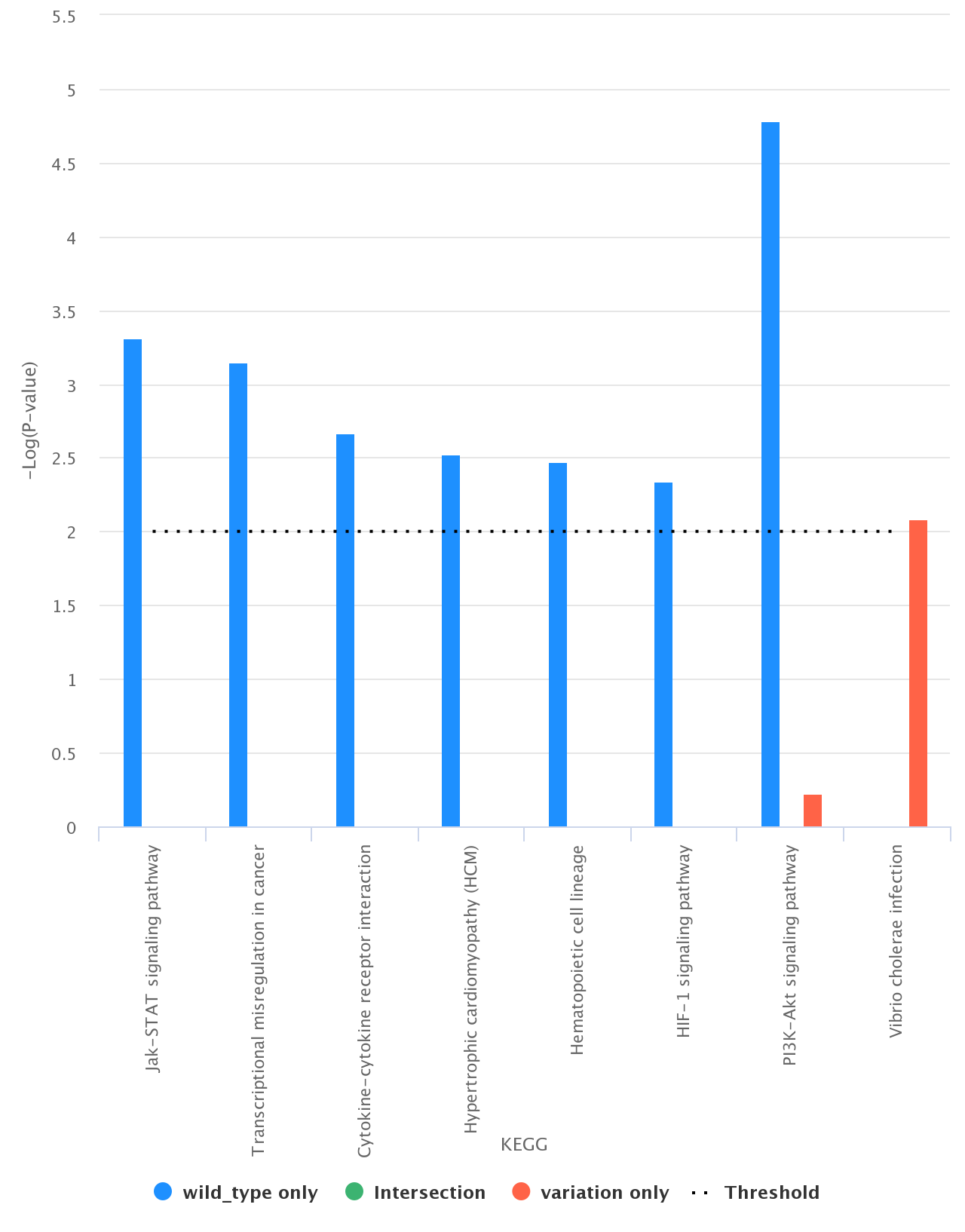
Histograms can be downloaded in different formats, such as PDF, PNG, JPEG, and SVG.
What is described above is valid for Reactome results as well.
Changelog
# Changelog All notable changes to this project will be documented in this section. The format is based on [Keep a Changelog](http://keepachangelog.com/en/1.0.0/) and this project adheres to [Semantic Versioning](http://semver.org/spec/v2.0.0.html). ## [Released] ## [1.2.1] - 2020-06-22 ### Fixed - Fix issues affecting the isoTar interface, where some input combinations of miRNA-pre-miRNA were discarded. ## [1.2] - 2020-03-26 ### Fixed - Fix issues affecting miRanda's predictions, setting -ge and -go values to the default values. Version v1.1 erroneously employed -ge 4 and -go 9). ## [1.1] - 2019-10-04 ### Fixed - Fix issues affecting the last nucleotide at 3'-end of miRNA in each RNAhybrid's prediction - Fix issues affecting miRmap's predictions for isomiRs and miRNAs with substitution. ### Changed - Fix typos in Homepage - Fix two PubMed IDs for A-to-I miRNA Database ## [1.0] - 2019-04-05 ### Added - Initial release